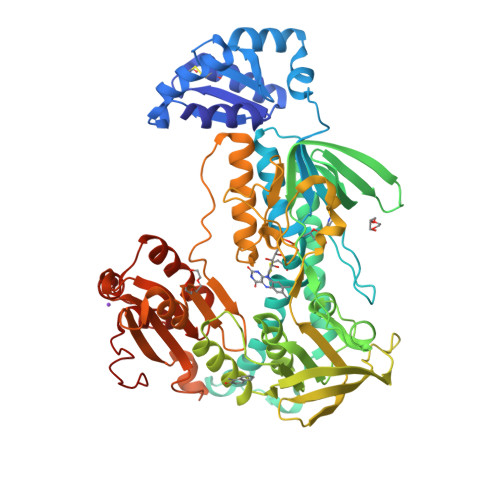
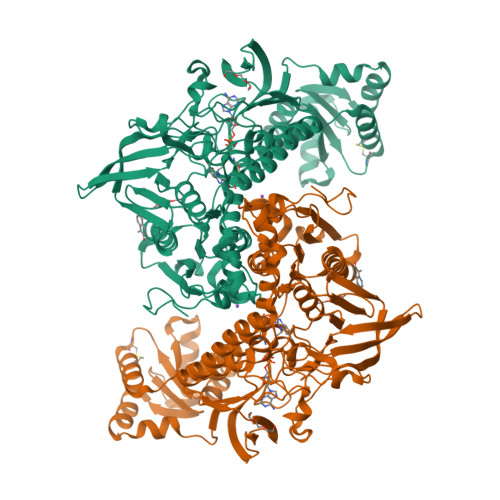
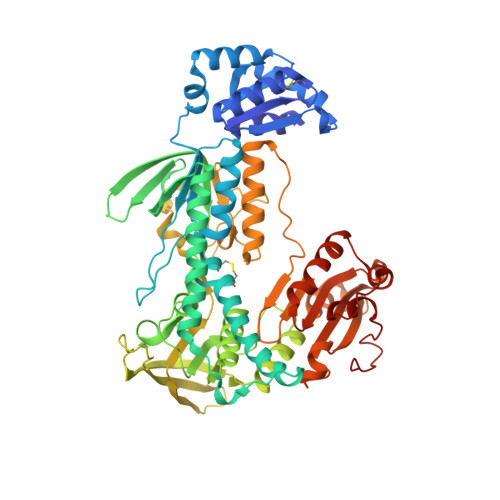
Ectopic suicide inhibition of thioredoxin glutathione reductase.
Silvestri, I., Lyu, H., Fata, F., Banta, P.R., Mattei, B., Ippoliti, R., Bellelli, A., Pitari, G., Ardini, M., Petukhova, V., Thatcher, G.R.J., Petukhov, P.A., Williams, D.L., Angelucci, F.(2020) Free Radic Biol Med 147: 200-211
- PubMed: 31870799
- DOI: https://doi.org/10.1016/j.freeradbiomed.2019.12.019
- Primary Citation of Related Structures:
6RTJ, 6RTM, 6RTO - PubMed Abstract:
Selective suicide inhibitors represent a seductively attractive approach for inactivation of therapeutically relevant enzymes since they are generally devoid of off-target toxicity in vivo. While most suicide inhibitors are converted to reactive species at enzyme active sites, theoretically bioactivation can also occur in ectopic (secondary) sites that have no known function. Here, we report an example of such an "ectopic suicide inhibition", an unprecedented bioactivation mechanism of a suicide inhibitor carried out by a non-catalytic site of thioredoxin glutathione reductase (TGR). TGR is a promising drug target to treat schistosomiasis, a devastating human parasitic disease. Utilizing hits selected from a high throughput screening campaign, time-resolved X-ray crystallography, molecular dynamics, mass spectrometry, molecular modeling, protein mutagenesis and functional studies, we find that 2-naphtholmethylamino derivatives bound to this novel ectopic site of Schistosoma mansoni (Sm)TGR are transformed to covalent modifiers and react with its mobile selenocysteine-containing C-terminal arm. In particular, one 2-naphtholmethylamino compound is able to specifically induce the pro-oxidant activity in the inhibited enzyme. Since some 2-naphtholmethylamino analogues show worm killing activity and the ectopic site is not conserved in human orthologues, a general approach to development of novel and selective anti-parasitic therapeutics against schistosoma is proposed.
Organizational Affiliation:
Dept. of Life, Health and Environmental Sciences, University of L'Aquila, Italy.