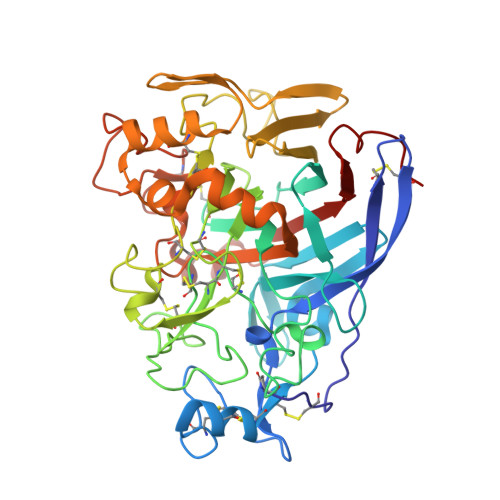
Enantioselective Binding of Propranolol and Analogues Thereof to Cellobiohydrolase Cel7A.
Hamark, C., Pendrill, R., Landstrom, J., Dotson Fagerstrom, A., Sandgren, M., Stahlberg, J., Widmalm, G.(2018) Chemistry 24: 17975-17985
- PubMed: 30255965
- DOI: https://doi.org/10.1002/chem.201803104
- Primary Citation of Related Structures:
6GRN - PubMed Abstract:
At the catalytic site for the hydrolysis of cellulose the enzyme cellobiohydrolase Cel7A binds the enantiomers of the adrenergic beta-blocker propranolol with different selectivity. Methyl-to-hydroxymethyl group modifications of propranolol, which result in higher affinity and improved selectivity, were herein studied by 1 H, 1 H and 1 H, 13 C scalar spin-spin coupling constants as well as utilizing the nuclear Overhauser effect (NOE) in conjunction with molecular dynamics simulations of the ligands per se, which showed the presence of all-antiperiplanar conformations, except for the one containing a vicinal oxygen-oxygen arrangement governed by the gauche effect. For the ligand-protein complexes investigated by NMR spectroscopy using, inter alia, transferred NOESY and saturation-transfer difference (STD) NMR experiments the S-isomers were shown to bind with a higher affinity and a conformation similar to that preferred in solution, in contrast to the R-isomer. The fact that the S-form of the propranolol enantiomer is pre-arranged for binding to the protein is also observed for a crystal structure of dihydroxy-(S)-propranolol and Cel7A presented herein. Whereas the binding of propranolol is entropy driven, the complexation with the dihydroxy analogue is anticipated to be favored also by an enthalpic term, such as for its enantiomer, that is, dihydroxy-(R)-propranolol, because hydrogen-bond donation replaces the corresponding bonding from hydroxyl groups in glucosyl residues of the natural substrate. In addition to a favorable entropy component, albeit lesser in magnitude, this represents an effect of enthalpy-to-entropy compensation in ligand-protein interactions.
Organizational Affiliation:
Department of Organic Chemistry, Arrhenius Laboratory, Stockholm University, 10691, Stockholm, Sweden.