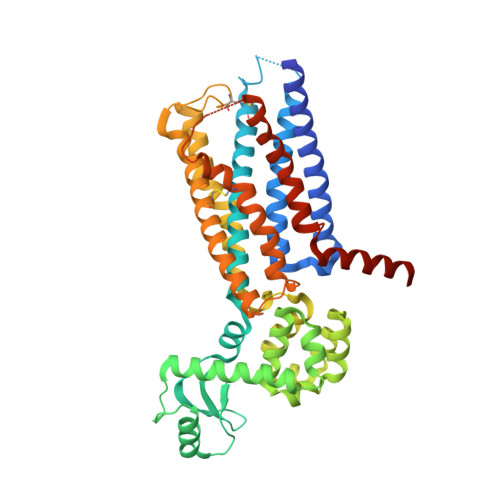
Mutagenesis facilitated crystallization of GLP-1R.
Xu, Y., Wang, Y., Wang, Y., Liu, K., Peng, Y., Yao, D., Tao, H., Liu, H., Song, G.(2019) IUCrJ 6: 996-1006
- PubMed: 31709055
- DOI: https://doi.org/10.1107/S2052252519013496
- Primary Citation of Related Structures:
6KJV, 6KK1, 6KK7 - PubMed Abstract:
The class B family of G-protein-coupled receptors (GPCRs) has long been a paradigm for peptide hormone recognition and signal transduction. One class B GPCR, the glucagon-like peptide-1 receptor (GLP-1R), has been considered as an anti-diabetes drug target and there are several peptidic drugs available for the treatment of this overwhelming disease. The previously determined structures of inactive GLP-1R in complex with two negative allosteric modulators include ten thermal-stabilizing mutations that were selected from a total of 98 designed mutations. Here we systematically summarize all 98 mutations we have tested and the results suggest that the mutagenesis strategy that strengthens inter-helical hydro-phobic interactions shows the highest success rate. We further investigate four back mutations by thermal-shift assay, crystallization and molecular dynamic simulations, and conclude that mutation I196 2.66b F increases thermal stability intrinsically and that mutation S271 4.47b A decreases crystal packing entropy extrinsically, while mutations S193 2.63b C and M233 3.36b C may be dispensable since these two cysteines are not di-sulfide-linked. Our results indicate intrinsic connections between different regions of GPCR transmembrane helices and the current data suggest a general mutagenesis principle for structural determination of GPCRs and other membrane proteins.
Organizational Affiliation:
Shanghai Key Laboratory of Regulatory Biology, Institute of Biomedical Sciences and School of Life Sciences, East China Normal University, Shanghai 200241, People's Republic of China.