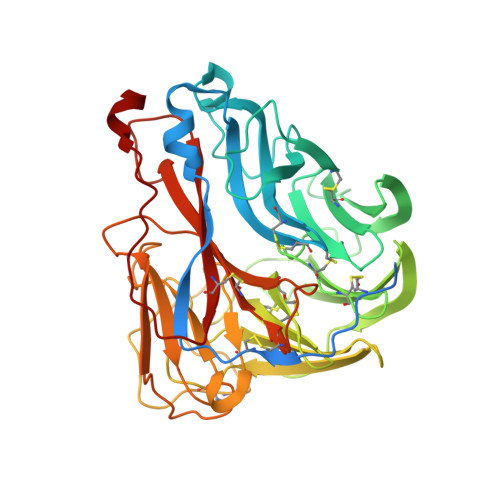
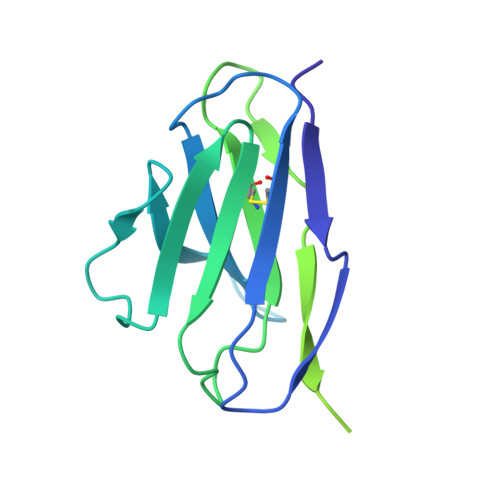
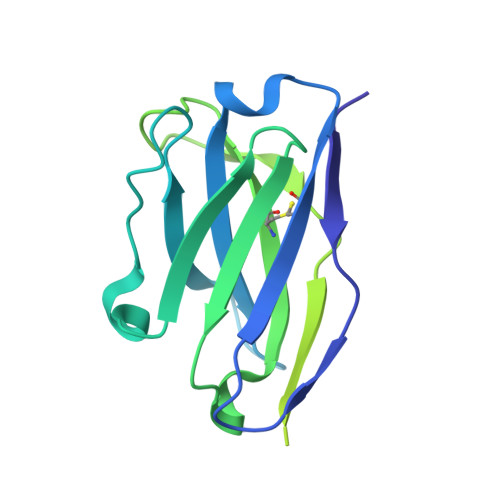
Structural Basis of Protection against H7N9 Influenza Virus by Human Anti-N9 Neuraminidase Antibodies.
Zhu, X., Turner, H.L., Lang, S., McBride, R., Bangaru, S., Gilchuk, I.M., Yu, W., Paulson, J.C., Crowe Jr., J.E., Ward, A.B., Wilson, I.A.(2019) Cell Host Microbe 26: 729
- PubMed: 31757767
- DOI: https://doi.org/10.1016/j.chom.2019.10.002
- Primary Citation of Related Structures:
6PZD, 6PZE, 6PZF, 6PZG, 6PZH, 6PZW, 6PZY, 6PZZ, 6U02 - PubMed Abstract:
Influenza virus neuraminidase (NA) is a major target for small-molecule antiviral drugs. Antibodies targeting the NA surface antigen could also inhibit virus entry and egress to provide host protection. However, our understanding of the nature and range of target epitopes is limited because of a lack of human antibody structures with influenza neuraminidase. Here, we describe crystal and cryogenic electron microscopy (cryo-EM) structures of NAs from human-infecting avian H7N9 viruses in complex with five human anti-N9 antibodies, systematically defining several antigenic sites and antibody epitope footprints. These antibodies either fully or partially block the NA active site or bind to epitopes distant from the active site while still showing neuraminidase inhibition. The inhibition of antibodies to NAs was further analyzed by glycan array and solution-based NA activity assays. Together, these structural studies provide insights into protection by anti-NA antibodies and templates for the development of NA-based influenza virus vaccines and therapeutics.
Organizational Affiliation:
Department of Integrative Structural and Computational Biology, The Scripps Research Institute, La Jolla, CA 92037, USA.