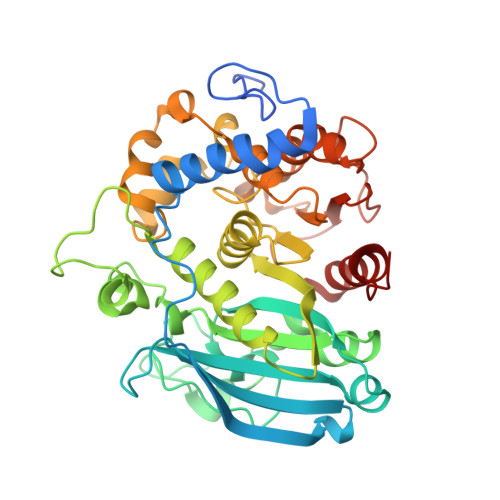
Structural and biochemical studies of the glucuronoyl esteraseOtCE15A illuminate its interaction with lignocellulosic components.
Mazurkewich, S., Poulsen, J.N., Lo Leggio, L., Larsbrink, J.(2019) J Biol Chem 294: 19978-19987
- PubMed: 31740581
- DOI: https://doi.org/10.1074/jbc.RA119.011435
- Primary Citation of Related Structures:
6SYR, 6SYU, 6SYV, 6SZ0, 6SZ4, 6SZO, 6T0E, 6T0I - PubMed Abstract:
Glucuronoyl esterases (GEs) catalyze the cleavage of ester linkages between lignin and glucuronic acid moieties on glucuronoxylan in plant biomass. As such, GEs represent promising biochemical tools in industrial processing of these recalcitrant resources. However, details on how GEs interact and catalyze degradation of their natural substrates are sparse, calling for thorough enzyme structure-function studies. Presented here is a structural and mechanistic investigation of the bacterial GE Ot CE15A. GEs belong to the carbohydrate esterase family 15 (CE15), which is in turn part of the larger α/β-hydrolase superfamily. GEs contain a Ser-His-Asp/Glu catalytic triad, but the location of the catalytic acid in GEs has been shown to be variable, and Ot CE15A possesses two putative catalytic acidic residues in the active site. Through site-directed mutagenesis, we demonstrate that these residues are functionally redundant, possibly indicating the evolutionary route toward new functionalities within the family. Structures determined with glucuronate, in both native and covalently bound intermediate states, and galacturonate provide insights into the catalytic mechanism of CE15. A structure of Ot CE15A with the glucuronoxylooligosaccharide 2 3 -(4- O -methyl-α-d-glucuronyl)-xylotriose (commonly referred to as XUX) shows that the enzyme can indeed interact with polysaccharides from the plant cell wall, and an additional structure with the disaccharide xylobiose revealed a surface binding site that could possibly indicate a recognition mechanism for long glucuronoxylan chains. Collectively, the results indicate that Ot CE15A, and likely most of the CE15 family, can utilize esters of glucuronoxylooligosaccharides and support the proposal that these enzymes work on lignin-carbohydrate complexes in plant biomass.
Organizational Affiliation:
Wallenberg Wood Science Center, Division of Industrial Biotechnology, Department of Biology and Biological Engineering, Chalmers University of Technology, SE-412 96 Gothenburg, Sweden.