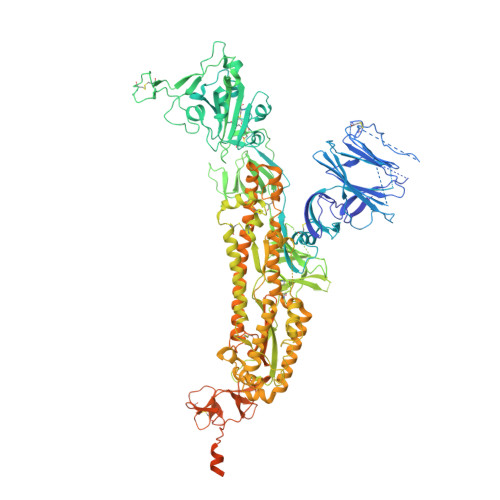
Neutralization of SARS-CoV-2 by Destruction of the Prefusion Spike.
Huo, J., Zhao, Y., Ren, J., Zhou, D., Duyvesteyn, H.M.E., Ginn, H.M., Carrique, L., Malinauskas, T., Ruza, R.R., Shah, P.N.M., Tan, T.K., Rijal, P., Coombes, N., Bewley, K.R., Tree, J.A., Radecke, J., Paterson, N.G., Supasa, P., Mongkolsapaya, J., Screaton, G.R., Carroll, M., Townsend, A., Fry, E.E., Owens, R.J., Stuart, D.I.(2020) Cell Host Microbe 28: 445-454.e6
- PubMed: 32585135
- DOI: https://doi.org/10.1016/j.chom.2020.06.010
- Primary Citation of Related Structures:
6YLA, 6YM0, 6YOR, 6Z97 - PubMed Abstract:
There are as yet no licensed therapeutics for the COVID-19 pandemic. The causal coronavirus (SARS-CoV-2) binds host cells via a trimeric spike whose receptor binding domain (RBD) recognizes angiotensin-converting enzyme 2, initiating conformational changes that drive membrane fusion. We find that the monoclonal antibody CR3022 binds the RBD tightly, neutralizing SARS-CoV-2, and report the crystal structure at 2.4 Å of the Fab/RBD complex. Some crystals are suitable for screening for entry-blocking inhibitors. The highly conserved, structure-stabilizing CR3022 epitope is inaccessible in the prefusion spike, suggesting that CR3022 binding facilitates conversion to the fusion-incompetent post-fusion state. Cryogenic electron microscopy (cryo-EM) analysis confirms that incubation of spike with CR3022 Fab leads to destruction of the prefusion trimer. Presentation of this cryptic epitope in an RBD-based vaccine might advantageously focus immune responses. Binders at this epitope could be useful therapeutically, possibly in synergy with an antibody that blocks receptor attachment.
Organizational Affiliation:
Division of Structural Biology, University of Oxford, The Wellcome Centre for Human Genetics, Headington, Oxford, OX3 7BN, UK; The Rosalind Franklin Institute, Harwell Campus, OX11 0FA, UK; Protein Production UK, Research Complex at Harwell, Harwell Science & Innovation Campus, Didcot, OX11 0FA, UK.