Structural Basis of a Human Neutralizing Antibody Specific to the SARS-CoV-2 Spike Protein Receptor-Binding Domain.
Yang, M., Li, J., Huang, Z., Li, H., Wang, Y., Wang, X., Kang, S., Huang, X., Wu, C., Liu, T., Jia, Z., Liang, J., Yuan, X., He, S., Chen, X., Zhou, Z., Chen, Q., Liu, S., Li, J., Zheng, H., Liu, X., Li, K., Yao, X., Lang, B., Liu, L., Liao, H.X., Chen, S.(2021) Microbiol Spectr 9: e0135221-e0135221
- PubMed: 34643438
- DOI: https://doi.org/10.1128/Spectrum.01352-21
- Primary Citation of Related Structures:
7E3O - PubMed Abstract:
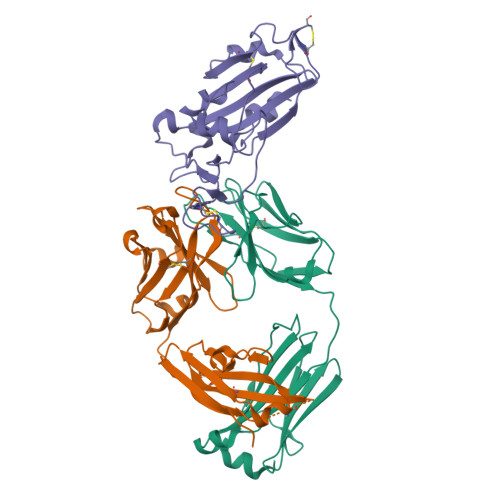
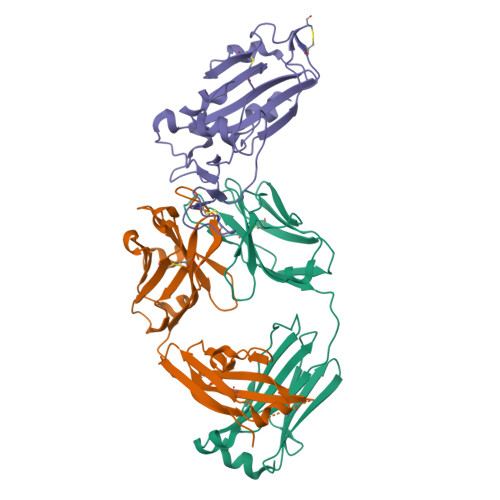
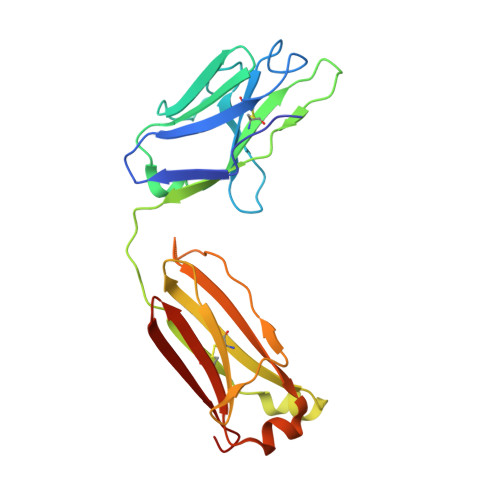
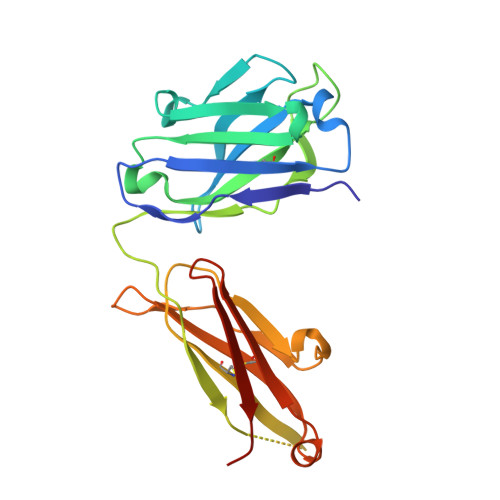
The emerging new lineages of severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) have marked a new phase of coronavirus disease 2019 (COVID-19). Understanding the recognition mechanisms of potent neutralizing monoclonal antibodies (NAbs) against the spike protein is pivotal for developing new vaccines and antibody drugs. Here, we isolated several monoclonal antibodies (MAbs) against the SARS-CoV-2 spike protein receptor-binding domain (S-RBD) from the B cell receptor repertoires of a SARS-CoV-2 convalescent. Among these MAbs, the antibody nCoV617 demonstrates the most potent neutralizing activity against authentic SARS-CoV-2 infection, as well as prophylactic and therapeutic efficacies against the human angiotensin-converting enzyme 2 (ACE2) transgenic mouse model in vivo . The crystal structure of S-RBD in complex with nCoV617 reveals that nCoV617 mainly binds to the back of the "ridge" of RBD and shares limited binding residues with ACE2. Under the background of the S-trimer model, it potentially binds to both "up" and "down" conformations of S-RBD. In vitro mutagenesis assays show that mutant residues found in the emerging new lineage B.1.1.7 of SARS-CoV-2 do not affect nCoV617 binding to the S-RBD. These results provide a new human-sourced neutralizing antibody against the S-RBD and assist vaccine development. IMPORTANCE COVID-19 is a respiratory disease caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). The COVID-19 pandemic has posed a serious threat to global health and the economy, so it is necessary to find safe and effective antibody drugs and treatments. The receptor-binding domain (RBD) in the SARS-CoV-2 spike protein is responsible for binding to the angiotensin-converting enzyme 2 (ACE2) receptor. It contains a variety of dominant neutralizing epitopes and is an important antigen for the development of new coronavirus antibodies. The significance of our research lies in the determination of new epitopes, the discovery of antibodies against RBD, and the evaluation of the antibodies' neutralizing effect. The identified antibodies here may be drug candidates for the development of clinical interventions for SARS-CoV-2.
Organizational Affiliation:
Molecular Imaging Center, Guangdong Provincial Key Laboratory of Biomedical Imaging, The Fifth Affiliated Hospital, Sun Yat-sen Universitygrid.12981.33, Zhuhai, China.