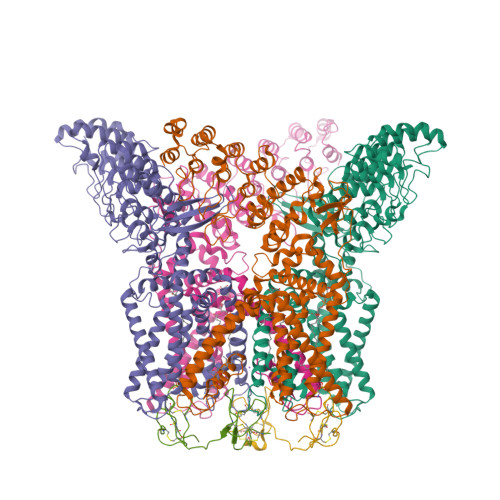
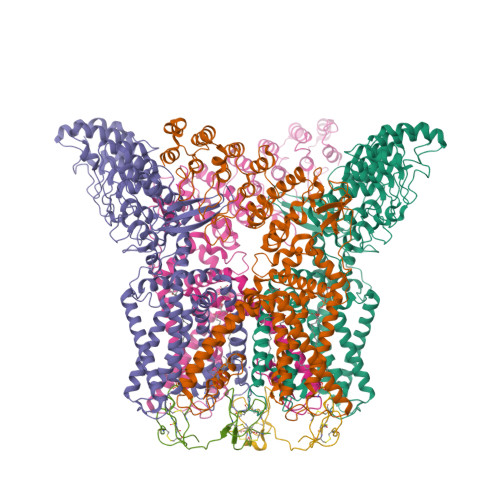
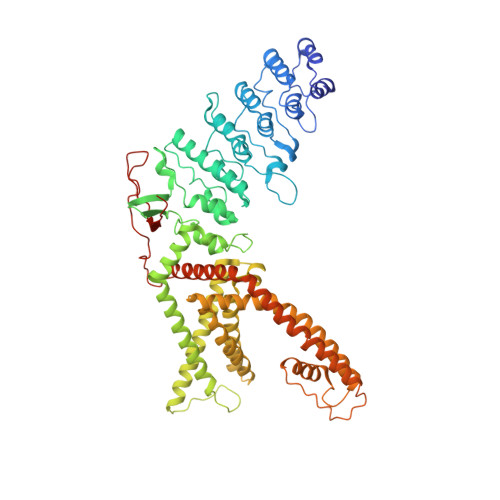
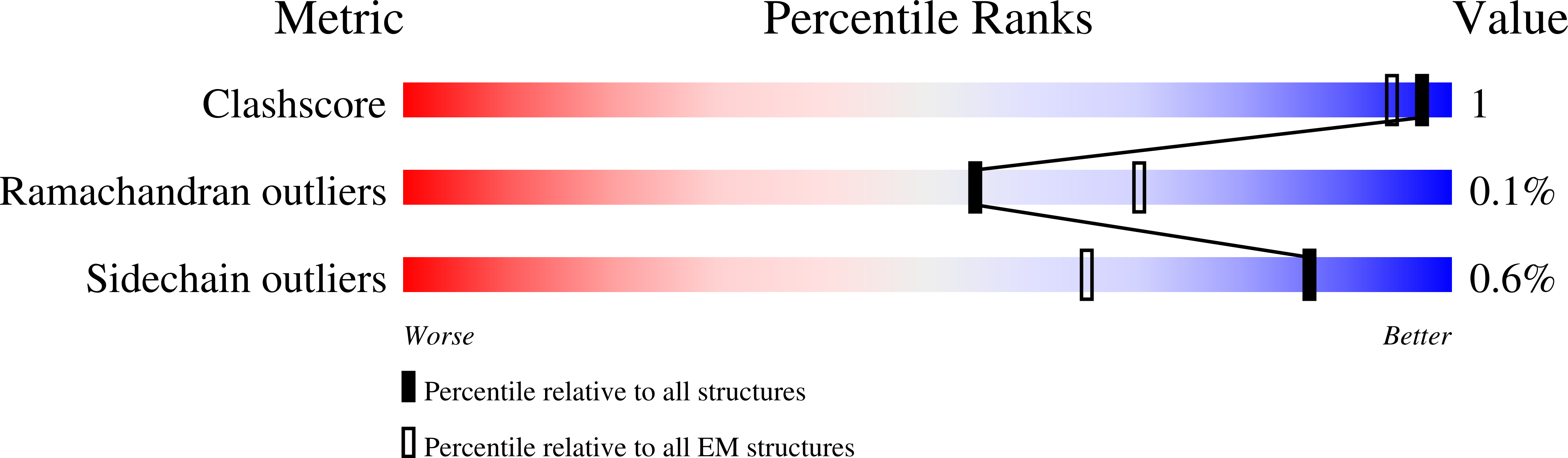Structural snapshots of TRPV1 reveal mechanism of polymodal functionality.
Zhang, K., Julius, D., Cheng, Y.(2021) Cell 184: 5138
- PubMed: 34496225
- DOI: https://doi.org/10.1016/j.cell.2021.08.012
- Primary Citation of Related Structures:
7L2H, 7L2I, 7L2J, 7L2K, 7L2L, 7L2M, 7L2N, 7L2O, 7L2P, 7L2R, 7L2S, 7L2T, 7L2U, 7L2V, 7L2W, 7L2X, 7MZ5, 7MZ6, 7MZ7, 7MZ9, 7MZA, 7MZB, 7MZC, 7MZD, 7MZE - PubMed Abstract:
Many transient receptor potential (TRP) channels respond to diverse stimuli and conditionally conduct small and large cations. Such functional plasticity is presumably enabled by a uniquely dynamic ion selectivity filter that is regulated by physiological agents. What is currently missing is a "photo series" of intermediate structural states that directly address this hypothesis and reveal specific mechanisms behind such dynamic channel regulation. Here, we exploit cryoelectron microscopy (cryo-EM) to visualize conformational transitions of the capsaicin receptor, TRPV1, as a model to understand how dynamic transitions of the selectivity filter in response to algogenic agents, including protons, vanilloid agonists, and peptide toxins, permit permeation by small and large organic cations. These structures also reveal mechanisms governing ligand binding substates, as well as allosteric coupling between key sites that are proximal to the selectivity filter and cytoplasmic gate. These insights suggest a general framework for understanding how TRP channels function as polymodal signal integrators.
Organizational Affiliation:
Department of Biochemistry and Biophysics, University of California, San Francisco, San Francisco, CA, USA.