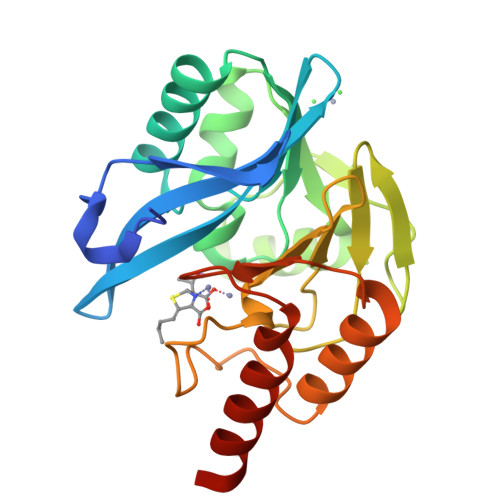
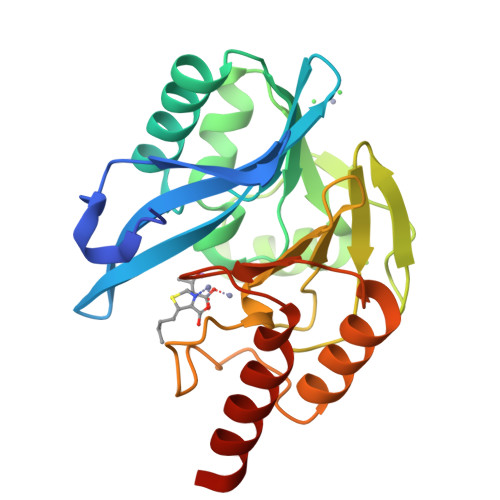
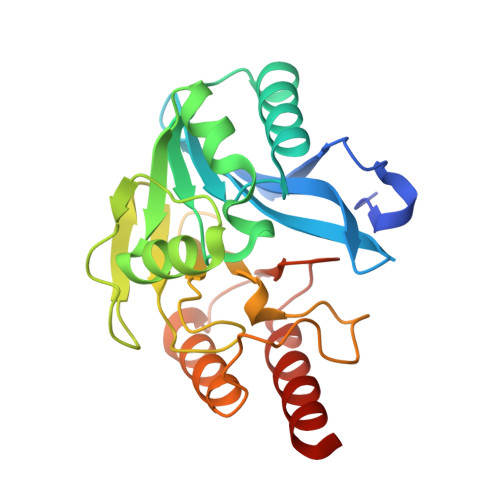
Faropenem reacts with serine and metallo-beta-lactamases to give multiple products.
Lucic, A., Hinchliffe, P., Malla, T.R., Tooke, C.L., Brem, J., Calvopina, K., Lohans, C.T., Rabe, P., McDonough, M.A., Armistead, T., Orville, A.M., Spencer, J., Schofield, C.J.(2021) Eur J Med Chem 215: 113257-113257
- PubMed: 33618159
- DOI: https://doi.org/10.1016/j.ejmech.2021.113257
- Primary Citation of Related Structures:
7A5Z, 7A60, 7A61, 7A63 - PubMed Abstract:
Penems have demonstrated potential as antibacterials and β-lactamase inhibitors; however, their clinical use has been limited, especially in comparison with the structurally related carbapenems. Faropenem is an orally active antibiotic with a C-2 tetrahydrofuran (THF) ring, which is resistant to hydrolysis by some β-lactamases. We report studies on the reactions of faropenem with carbapenem-hydrolysing β-lactamases, focusing on the class A serine β-lactamase KPC-2 and the metallo β-lactamases (MBLs) VIM-2 (a subclass B1 MBL) and L1 (a B3 MBL). Kinetic studies show that faropenem is a substrate for all three β-lactamases, though it is less efficiently hydrolysed by KPC-2. Crystallographic analyses on faropenem-derived complexes reveal opening of the β-lactam ring with formation of an imine with KPC-2, VIM-2, and L1. In the cases of the KPC-2 and VIM-2 structures, the THF ring is opened to give an alkene, but with L1 the THF ring remains intact. Solution state studies, employing NMR, were performed on L1, KPC-2, VIM-2, VIM-1, NDM-1, OXA-23, OXA-10, and OXA-48. The solution results reveal, in all cases, formation of imine products in which the THF ring is opened; formation of a THF ring-closed imine product was only observed with VIM-1 and VIM-2. An enamine product with a closed THF ring was also observed in all cases, at varying levels. Combined with previous reports, the results exemplify the potential for different outcomes in the reactions of penems with MBLs and SBLs and imply further structure-activity relationship studies are worthwhile to optimise the interactions of penems with β-lactamases. They also exemplify how crystal structures of β-lactamase substrate/inhibitor complexes do not always reflect reaction outcomes in solution.
Organizational Affiliation:
Chemistry Research Laboratory, The Department of Chemistry and the Ineos Oxford Institute for Antimicrobial Research, 12 Mansfield Road, Oxford, OX1 3TA, United Kingdom.