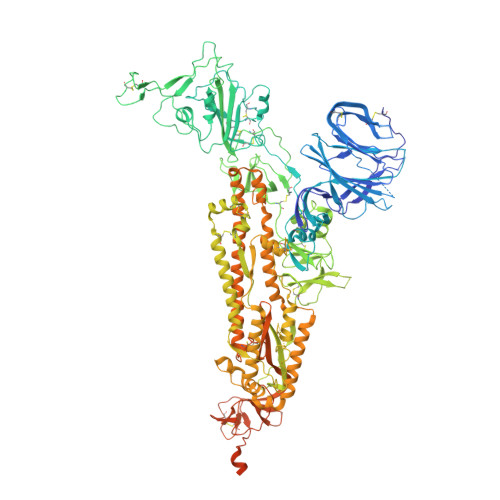
Structure and binding properties of Pangolin-CoV spike glycoprotein inform the evolution of SARS-CoV-2.
Wrobel, A.G., Benton, D.J., Xu, P., Calder, L.J., Borg, A., Roustan, C., Martin, S.R., Rosenthal, P.B., Skehel, J.J., Gamblin, S.J.(2021) Nat Commun 12: 837-837
- PubMed: 33547281
- DOI: https://doi.org/10.1038/s41467-021-21006-9
- Primary Citation of Related Structures:
7BBH - PubMed Abstract:
Coronaviruses of bats and pangolins have been implicated in the origin and evolution of the pandemic SARS-CoV-2. We show that spikes from Guangdong Pangolin-CoVs, closely related to SARS-CoV-2, bind strongly to human and pangolin ACE2 receptors. We also report the cryo-EM structure of a Pangolin-CoV spike protein and show it adopts a fully-closed conformation and that, aside from the Receptor-Binding Domain, it resembles the spike of a bat coronavirus RaTG13 more than that of SARS-CoV-2.
Organizational Affiliation:
Structural Biology of Disease Processes Laboratory, Francis Crick Institute, NW1 1AT, London, UK. antoni.wrobel@crick.ac.uk.