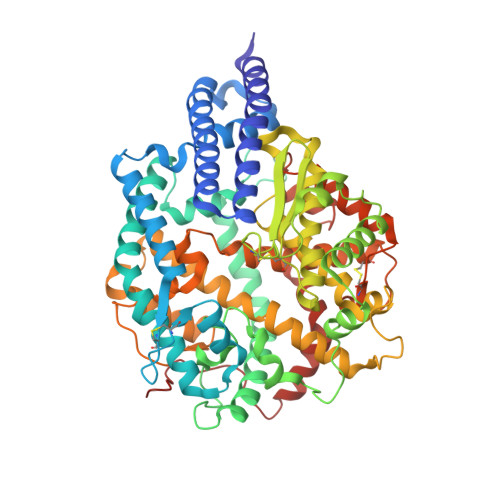
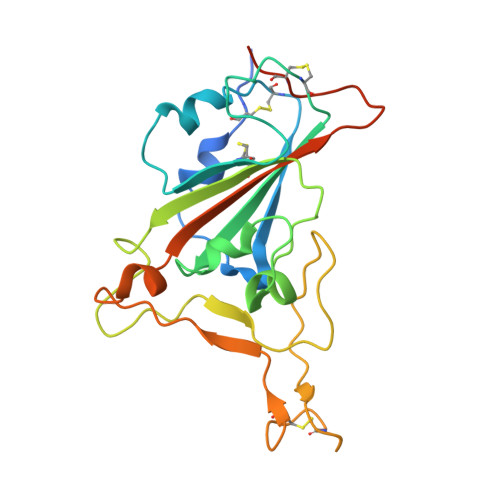
Binding and molecular basis of the bat coronavirus RaTG13 virus to ACE2 in humans and other species.
Liu, K., Pan, X., Li, L., Yu, F., Zheng, A., Du, P., Han, P., Meng, Y., Zhang, Y., Wu, L., Chen, Q., Song, C., Jia, Y., Niu, S., Lu, D., Qiao, C., Chen, Z., Ma, D., Ma, X., Tan, S., Zhao, X., Qi, J., Gao, G.F., Wang, Q.(2021) Cell 184: 3438-3451.e10
- PubMed: 34139177
- DOI: https://doi.org/10.1016/j.cell.2021.05.031
- Primary Citation of Related Structures:
7DRV - PubMed Abstract:
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has been spreading worldwide, causing a global pandemic. Bat-origin RaTG13 is currently the most phylogenetically related virus. Here we obtained the complex structure of the RaTG13 receptor binding domain (RBD) with human ACE2 (hACE2) and evaluated binding of RaTG13 RBD to 24 additional ACE2 orthologs. By substituting residues in the RaTG13 RBD with their counterparts in the SARS-CoV-2 RBD, we found that residue 501, the major position found in variants of concern (VOCs) 501Y.V1/V2/V3, plays a key role in determining the potential host range of RaTG13. We also found that SARS-CoV-2 could induce strong cross-reactive antibodies to RaTG13 and identified a SARS-CoV-2 monoclonal antibody (mAb), CB6, that could cross-neutralize RaTG13 pseudovirus. These results elucidate the receptor binding and host adaption mechanisms of RaTG13 and emphasize the importance of continuous surveillance of coronaviruses (CoVs) carried by animal reservoirs to prevent another spillover of CoVs.
Organizational Affiliation:
CAS Key Laboratory of Pathogenic Microbiology and Immunology, Institute of Microbiology, Chinese Academy of Sciences, Beijing 100101, China.