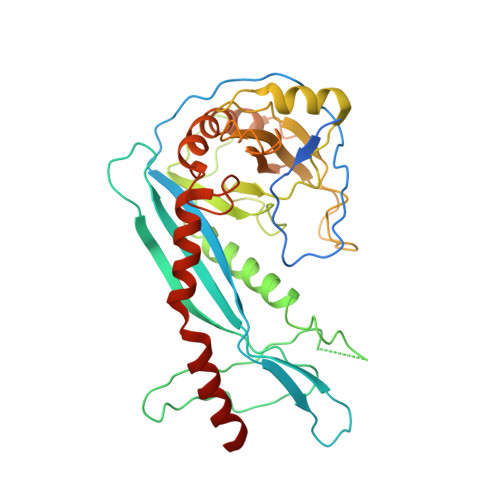
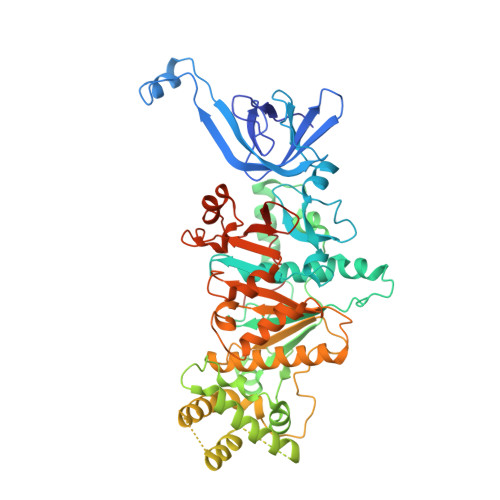
Mechanisms of helicase activated DNA end resection in bacteria.
Xu, Y., Xu, L., Qin, C., Wang, L., Guo, J., Hua, Y., Zhao, Y.(2022) Structure 30: 1298-1306.e3
- PubMed: 35841886
- DOI: https://doi.org/10.1016/j.str.2022.06.005
- Primary Citation of Related Structures:
7F6D - PubMed Abstract:
DNA end resection mediated by the coordinated action of nuclease and helicase is a crucial step in initiating homologous recombination. The end-resection apparatus NurA nuclease and HerA helicase are present in both archaea and bacteria. Here, we report the cryo-electron microscopy structure of a bacterial HerA-NurA complex from Deinococcus radiodurans. The structure reveals a barrel-like hexameric HerA and a distinctive NurA dimer subcomplex, which has a unique extended N-terminal region (ENR) involved in bacterial NurA dimerization and activation. In addition to the long protruding linking loop and the C-terminal α helix of NurA, the flexible ENR is close to the HerA-NurA interface and divides the central channel of the DrNurA dimer into two halves, suggesting a possible mechanism of DNA end processing. In summary, this work provides new insights into the structure, assembly, and activation mechanisms of bacterial DNA end resection mediated by a minimal end-resection apparatus.
Organizational Affiliation:
MOE Key Laboratory of Biosystems Homeostasis & Protection, Institute of Biophysics, College of Life Sciences, Zhejiang University, Hangzhou, China.