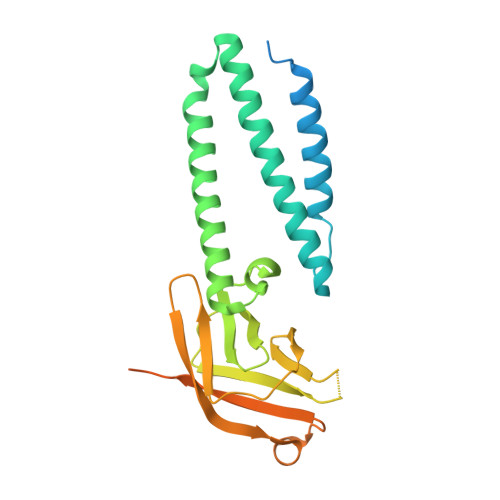
Cryo-EM structure of SARS-CoV-2 ORF3a in lipid nanodiscs.
Kern, D.M., Sorum, B., Mali, S.S., Hoel, C.M., Sridharan, S., Remis, J.P., Toso, D.B., Kotecha, A., Bautista, D.M., Brohawn, S.G.(2021) Nat Struct Mol Biol 28: 573-582
- PubMed: 34158638
- DOI: https://doi.org/10.1038/s41594-021-00619-0
- Primary Citation of Related Structures:
6XDC, 7KJR - PubMed Abstract:
SARS-CoV-2 ORF3a is a putative viral ion channel implicated in autophagy inhibition, inflammasome activation and apoptosis. 3a protein and anti-3a antibodies are found in infected patient tissues and plasma. Deletion of 3a in SARS-CoV-1 reduces viral titer and morbidity in mice, suggesting it could be an effective target for vaccines or therapeutics. Here, we present structures of SARS-CoV-2 3a determined by cryo-EM to 2.1-Å resolution. 3a adopts a new fold with a polar cavity that opens to the cytosol and membrane through separate water- and lipid-filled openings. Hydrophilic grooves along outer helices could form ion-conduction paths. Using electrophysiology and fluorescent ion imaging of 3a-reconstituted liposomes, we observe Ca 2+ -permeable, nonselective cation channel activity, identify mutations that alter ion permeability and discover polycationic inhibitors of 3a activity. 3a-like proteins are found across coronavirus lineages that infect bats and humans, suggesting that 3a-targeted approaches could treat COVID-19 and other coronavirus diseases.
Organizational Affiliation:
Department of Molecular & Cell Biology, University of California Berkeley, Berkeley, CA, USA.