Paired heavy- and light-chain signatures contribute to potent SARS-CoV-2 neutralization in public antibody responses.
Banach, B.B., Cerutti, G., Fahad, A.S., Shen, C.H., Oliveira De Souza, M., Katsamba, P.S., Tsybovsky, Y., Wang, P., Nair, M.S., Huang, Y., Francino-Urdaniz, I.M., Steiner, P.J., Gutierrez-Gonzalez, M., Liu, L., Lopez Acevedo, S.N., Nazzari, A.F., Wolfe, J.R., Luo, Y., Olia, A.S., Teng, I.T., Yu, J., Zhou, T., Reddem, E.R., Bimela, J., Pan, X., Madan, B., Laflin, A.D., Nimrania, R., Yuen, K.Y., Whitehead, T.A., Ho, D.D., Kwong, P.D., Shapiro, L., DeKosky, B.J.(2021) Cell Rep 37: 109771-109771
- PubMed: 34587480
- DOI: https://doi.org/10.1016/j.celrep.2021.109771
- Primary Citation of Related Structures:
7KS9 - PubMed Abstract:
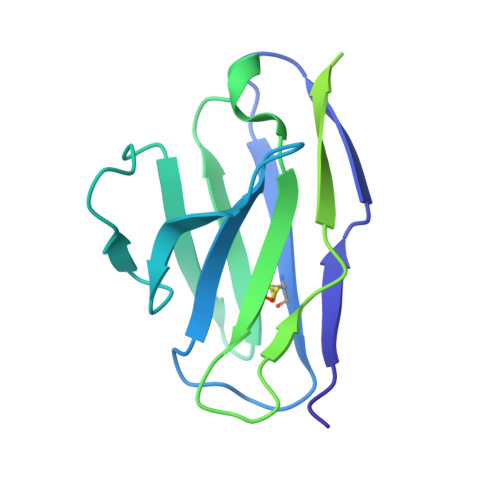
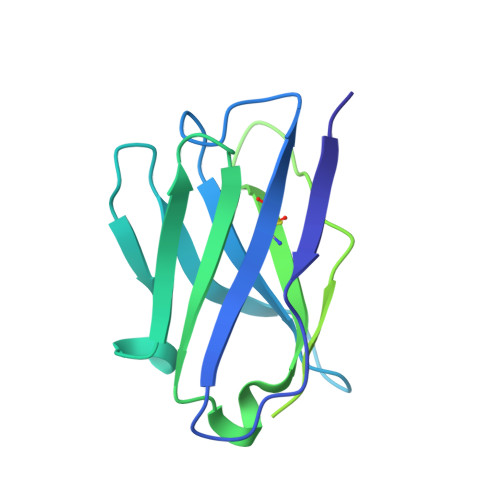
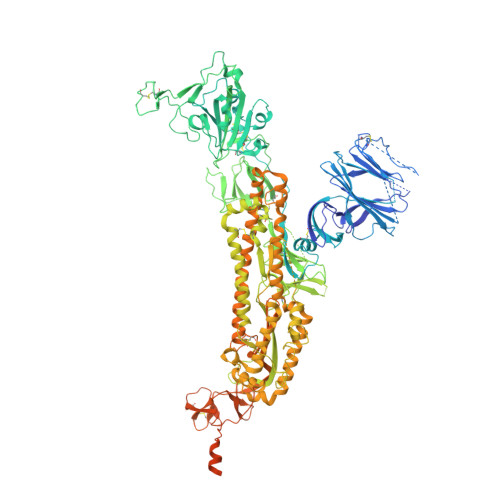
Understanding mechanisms of protective antibody recognition can inform vaccine and therapeutic strategies against SARS-CoV-2. We report a monoclonal antibody, 910-30, targeting the SARS-CoV-2 receptor-binding site for ACE2 as a member of a public antibody response encoded by IGHV3-53/IGHV3-66 genes. Sequence and structural analyses of 910-30 and related antibodies explore how class recognition features correlate with SARS-CoV-2 neutralization. Cryo-EM structures of 910-30 bound to the SARS-CoV-2 spike trimer reveal binding interactions and its ability to disassemble spike. Despite heavy-chain sequence similarity, biophysical analyses of IGHV3-53/3-66-encoded antibodies highlight the importance of native heavy:light pairings for ACE2-binding competition and SARS-CoV-2 neutralization. We develop paired heavy:light class sequence signatures and determine antibody precursor prevalence to be ∼1 in 44,000 human B cells, consistent with public antibody identification in several convalescent COVID-19 patients. These class signatures reveal genetic, structural, and functional immune features that are helpful in accelerating antibody-based medical interventions for SARS-CoV-2.
Organizational Affiliation:
Bioengineering Graduate Program, University of Kansas, Lawrence, KS 66045, USA.