Structural Basis for SARS-CoV-2 Nucleocapsid Protein Recognition by Single-Domain Antibodies.
Ye, Q., Lu, S., Corbett, K.D.(2021) Front Immunol 12: 719037-719037
- PubMed: 34381460
- DOI: https://doi.org/10.3389/fimmu.2021.719037
- Primary Citation of Related Structures:
7N0I, 7N0R, 7R98 - PubMed Abstract:
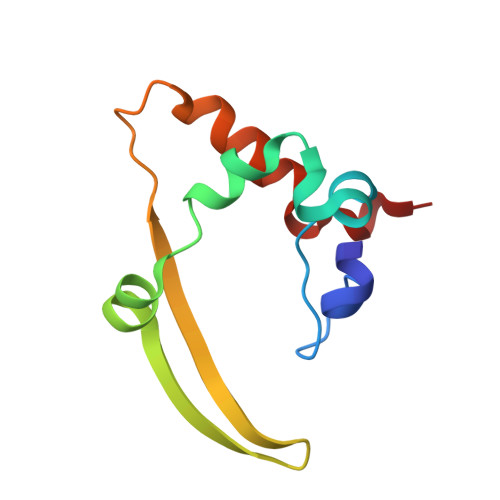
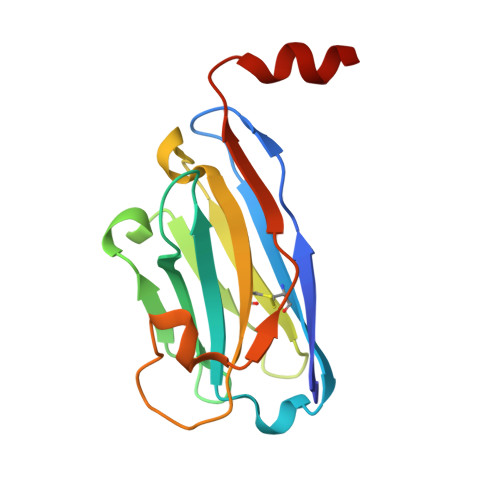
The COVID-19 pandemic, caused by the coronavirus SARS-CoV-2, is the most severe public health event of the twenty-first century. While effective vaccines against SARS-CoV-2 have been developed, there remains an urgent need for diagnostics to quickly and accurately detect infections. Antigen tests, particularly those that detect the abundant SARS-CoV-2 Nucleocapsid protein, are a proven method for detecting active SARS-CoV-2 infections. Here we report high-resolution crystal structures of three llama-derived single-domain antibodies that bind the SARS-CoV-2 Nucleocapsid protein with high affinity. Each antibody recognizes a specific folded domain of the protein, with two antibodies recognizing the N-terminal RNA binding domain and one recognizing the C-terminal dimerization domain. The two antibodies that recognize the RNA binding domain affect both RNA binding affinity and RNA-mediated phase separation of the Nucleocapsid protein. All three antibodies recognize highly conserved surfaces on the Nucleocapsid protein, suggesting that they could be used to develop affordable diagnostic tests to detect all circulating SARS-CoV-2 variants.
Organizational Affiliation:
Department of Cellular & Molecular Medicine, University of California San Diego, La Jolla, CA, United States.