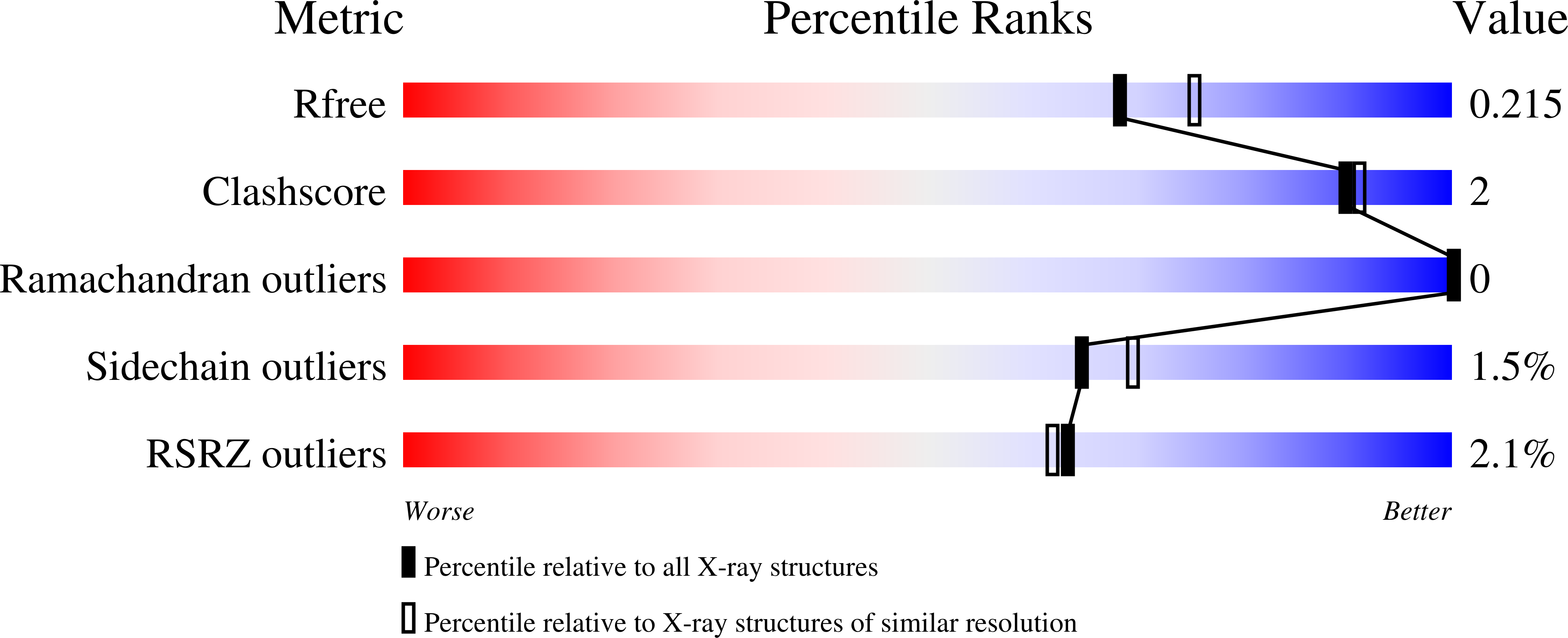
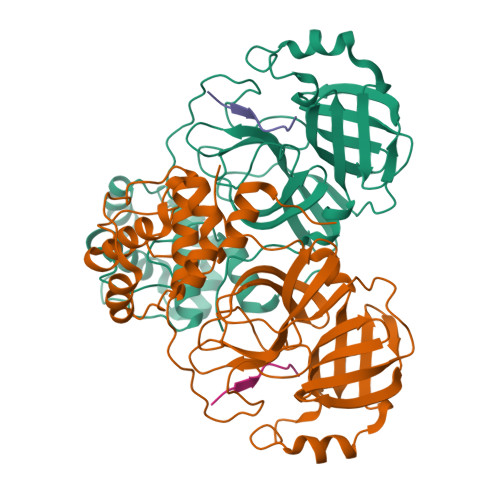
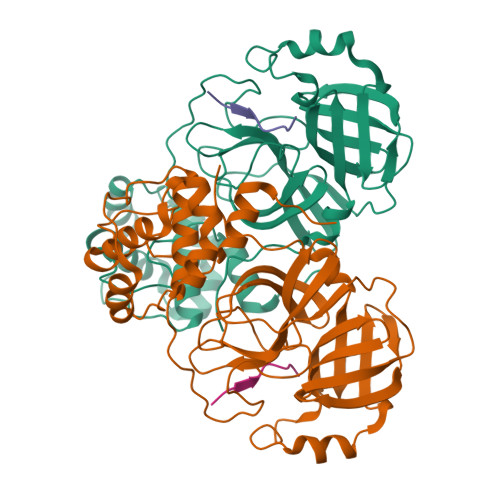
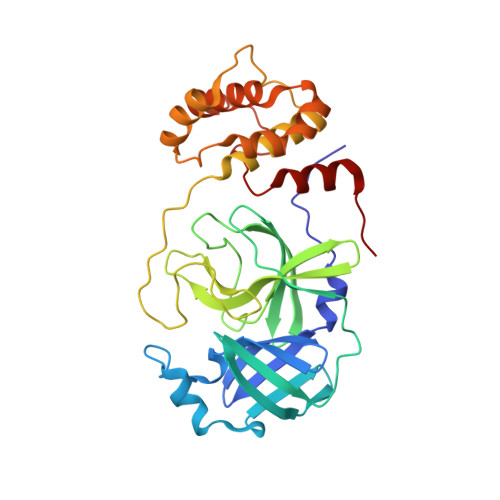
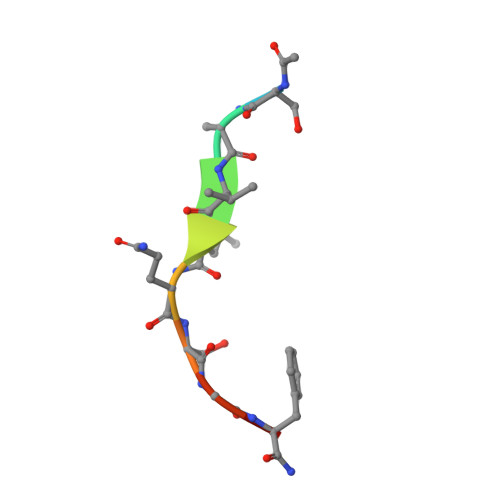
Michaelis-like complex of SARS-CoV-2 main protease visualized by room-temperature X-ray crystallography.
Kneller, D.W., Zhang, Q., Coates, L., Louis, J.M., Kovalevsky, A.(2021) IUCrJ 8: 973-979
- PubMed: 34804549
- DOI: https://doi.org/10.1107/S2052252521010113
- Primary Citation of Related Structures:
7N89 - PubMed Abstract:
SARS-CoV-2 emerged at the end of 2019 to cause an unprecedented pandemic of the deadly respiratory disease COVID-19 that continues to date. The viral main protease (M pro ) is essential for SARS-CoV-2 replication and is therefore an important drug target. Understanding the catalytic mechanism of M pro , a cysteine protease with a catalytic site comprising the noncanonical Cys145-His41 dyad, can help in guiding drug design. Here, a 2.0 Å resolution room-temperature X-ray crystal structure is reported of a Michaelis-like complex of M pro harboring a single inactivating mutation C145A bound to the octapeptide Ac-SAVLQSGF-CONH 2 corresponding to the nsp4/nsp5 autocleavage site. The peptide substrate is unambiguously defined in subsites S5 to S3' by strong electron density. Superposition of the Michaelis-like complex with the neutron structure of substrate-free M pro demonstrates that the catalytic site is inherently pre-organized for catalysis prior to substrate binding. Induced fit to the substrate is driven by P1 Gln binding in the predetermined subsite S1 and rearrangement of subsite S2 to accommodate P2 Leu. The Michaelis-like complex structure is ideal for in silico modeling of the SARS-CoV-2 M pro catalytic mechanism.
Organizational Affiliation:
Neutron Scattering Division, Oak Ridge National Laboratory, 1 Bethel Valley Road, Oak Ridge, TN 37831, USA.