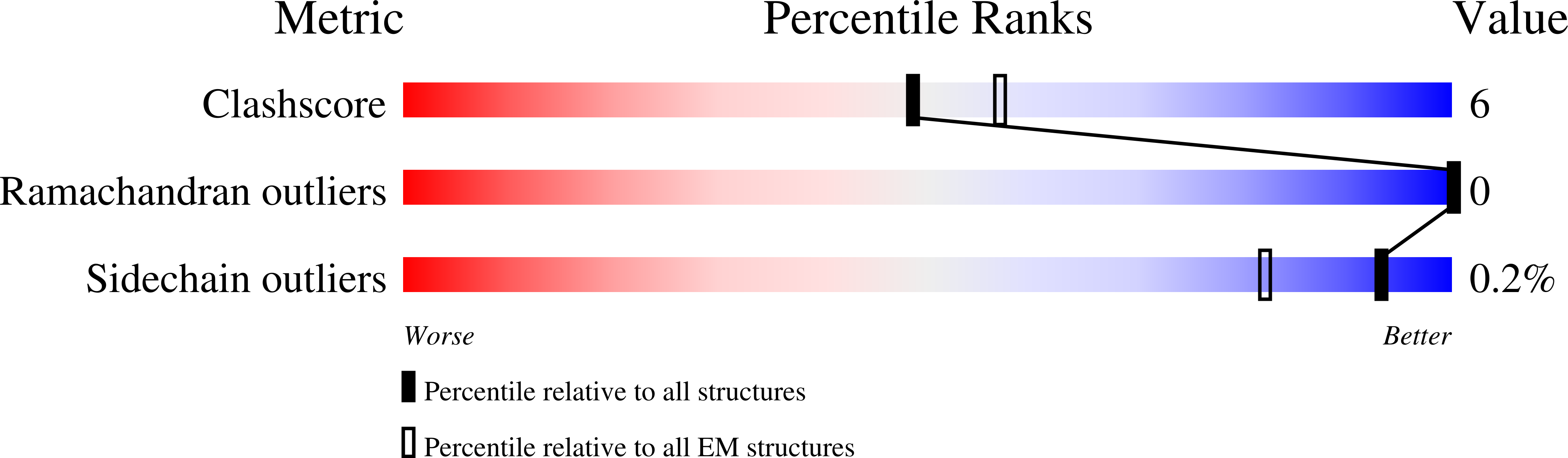Immunization with synthetic SARS-CoV-2 S glycoprotein virus-like particles protects macaques from infection.
Sulbaran, G., Maisonnasse, P., Amen, A., Effantin, G., Guilligay, D., Dereuddre-Bosquet, N., Burger, J.A., Poniman, M., Grobben, M., Buisson, M., Dergan Dylon, S., Naninck, T., Lemaitre, J., Gros, W., Gallouet, A.S., Marlin, R., Bouillier, C., Contreras, V., Relouzat, F., Fenel, D., Thepaut, M., Bally, I., Thielens, N., Fieschi, F., Schoehn, G., van der Werf, S., van Gils, M.J., Sanders, R.W., Poignard, P., Le Grand, R., Weissenhorn, W.(2022) Cell Rep Med 3: 100528-100528
- PubMed: 35233549
- DOI: https://doi.org/10.1016/j.xcrm.2022.100528
- Primary Citation of Related Structures:
7Q1Z - PubMed Abstract:
The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) pandemic has caused an ongoing global health crisis. Here, we present as a vaccine candidate synthetic SARS-CoV-2 spike (S) glycoprotein-coated lipid vesicles that resemble virus-like particles. Soluble S glycoprotein trimer stabilization by formaldehyde cross-linking introduces two major inter-protomer cross-links that keep all receptor-binding domains in the "down" conformation. Immunization of cynomolgus macaques with S coated onto lipid vesicles (S-LVs) induces high antibody titers with potent neutralizing activity against the vaccine strain, Alpha, Beta, and Gamma variants as well as T helper (Th)1 CD4 + -biased T cell responses. Although anti-receptor-binding domain (RBD)-specific antibody responses are initially predominant, the third immunization boosts significant non-RBD antibody titers. Challenging vaccinated animals with SARS-CoV-2 shows a complete protection through sterilizing immunity, which correlates with the presence of nasopharyngeal anti-S immunoglobulin G (IgG) and IgA titers. Thus, the S-LV approach is an efficient and safe vaccine candidate based on a proven classical approach for further development and clinical testing.
Organizational Affiliation:
Univ. Grenoble Alpes, CEA, CNRS, Institut de Biologie Structurale (IBS), Grenoble, France.