Potent human broadly SARS-CoV-2-neutralizing IgA and IgG antibodies effective against Omicron BA.1 and BA.2.
Planchais, C., Fernandez, I., Bruel, T., de Melo, G.D., Prot, M., Beretta, M., Guardado-Calvo, P., Dufloo, J., Molinos-Albert, L.M., Backovic, M., Chiaravalli, J., Giraud, E., Vesin, B., Conquet, L., Grzelak, L., Planas, D., Staropoli, I., Guivel-Benhassine, F., Hieu, T., Boulle, M., Cervantes-Gonzalez, M., Ungeheuer, M.N., Charneau, P., van der Werf, S., Agou, F., Dimitrov, J.D., Simon-Loriere, E., Bourhy, H., Montagutelli, X., Rey, F.A., Schwartz, O., Mouquet, H.(2022) J Exp Med 219
- PubMed: 35704748
- DOI: https://doi.org/10.1084/jem.20220638
- Primary Citation of Related Structures:
7QEZ, 7QF0, 7QF1 - PubMed Abstract:
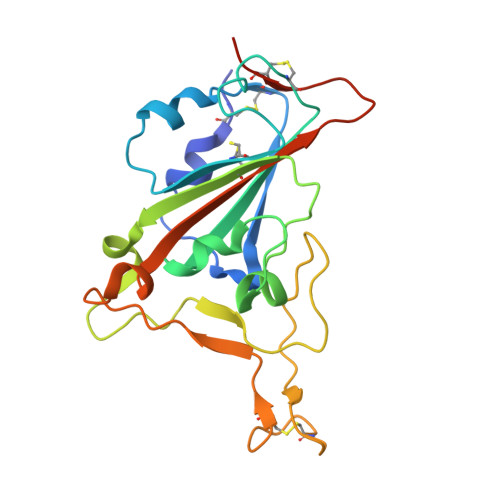
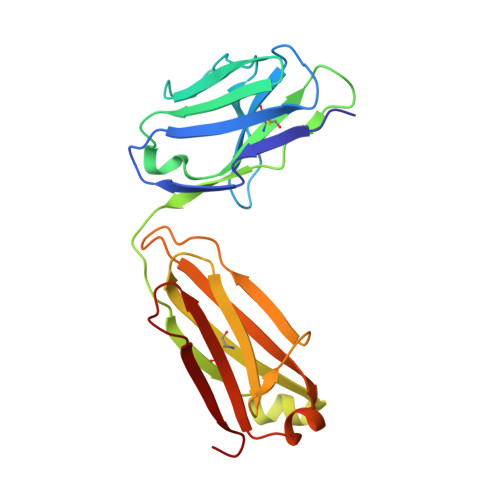
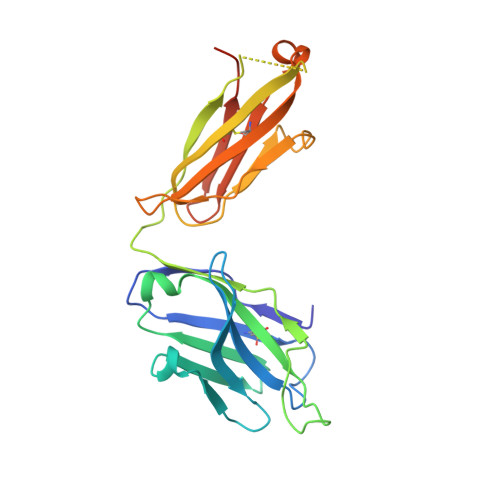
Memory B-cell and antibody responses to the SARS-CoV-2 spike protein contribute to long-term immune protection against severe COVID-19, which can also be prevented by antibody-based interventions. Here, wide SARS-CoV-2 immunoprofiling in Wuhan COVID-19 convalescents combining serological, cellular, and monoclonal antibody explorations revealed humoral immunity coordination. Detailed characterization of a hundred SARS-CoV-2 spike memory B-cell monoclonal antibodies uncovered diversity in their repertoire and antiviral functions. The latter were influenced by the targeted spike region with strong Fc-dependent effectors to the S2 subunit and potent neutralizers to the receptor-binding domain. Amongst those, Cv2.1169 and Cv2.3194 antibodies cross-neutralized SARS-CoV-2 variants of concern, including Omicron BA.1 and BA.2. Cv2.1169, isolated from a mucosa-derived IgA memory B cell demonstrated potency boost as IgA dimers and therapeutic efficacy as IgG antibodies in animal models. Structural data provided mechanistic clues to Cv2.1169 potency and breadth. Thus, potent broadly neutralizing IgA antibodies elicited in mucosal tissues can stem SARS-CoV-2 infection, and Cv2.1169 and Cv2.3194 are prime candidates for COVID-19 prevention and treatment.
Organizational Affiliation:
Institut Pasteur, Université Paris Cité, Laboratory of Humoral Immunology, Paris, France.