Structure of a Vaccine-Induced, Germline-Encoded Human Antibody Defines a Neutralizing Epitope on the SARS-CoV-2 Spike N-Terminal Domain.
Altomare, C.G., Adelsberg, D.C., Carreno, J.M., Sapse, I.A., Amanat, F., Ellebedy, A.H., Simon, V., Krammer, F., Bajic, G.(2022) mBio 13: e0358021-e0358021
- PubMed: 35467422
- DOI: https://doi.org/10.1128/mbio.03580-21
- Primary Citation of Related Structures:
7RBU, 7RBV - PubMed Abstract:
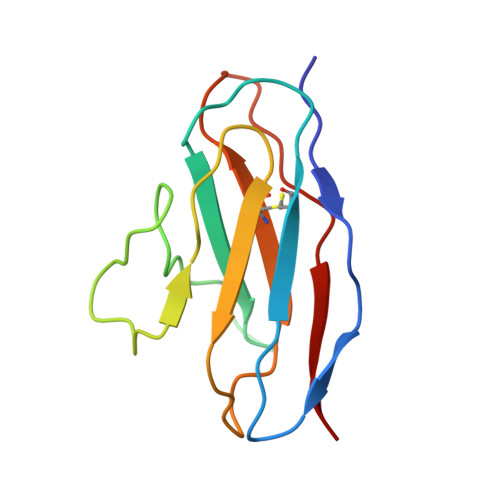
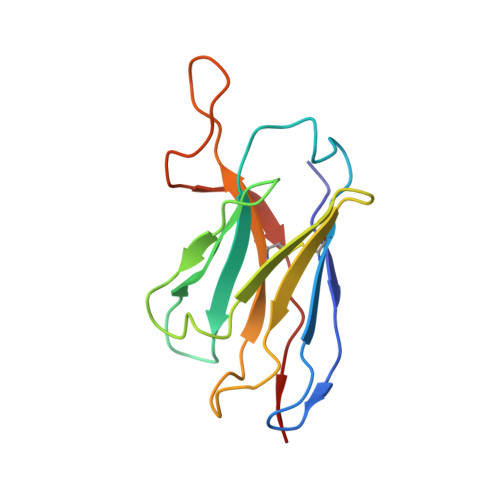
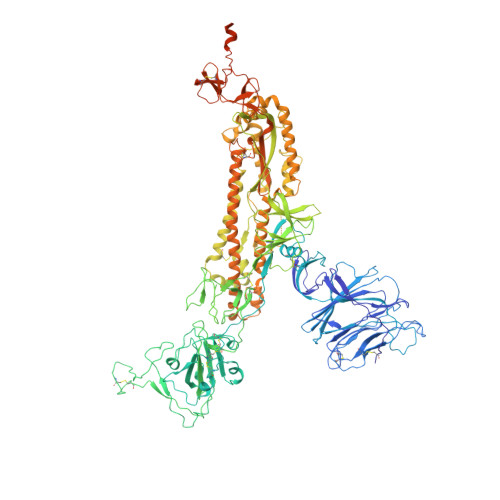
Structural characterization of infection- and vaccination-elicited antibodies in complex with antigen provides insight into the evolutionary arms race between the host and the pathogen and informs rational vaccine immunogen design. We isolated a germ line-encoded monoclonal antibody (mAb) from plasmablasts activated upon mRNA vaccination against severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and determined its structure in complex with the spike glycoprotein by electron cryomicroscopy (cryo-EM). We show that the mAb engages a previously uncharacterized neutralizing epitope on the spike N-terminal domain (NTD). The high-resolution structure reveals details of the intermolecular interactions and shows that the mAb inserts its heavy complementarity-determining region 3 (HCDR3) loop into a hydrophobic NTD cavity previously shown to bind a heme metabolite, biliverdin. We demonstrate direct competition with biliverdin and that, because of the conserved nature of the epitope, the mAb maintains binding to viral variants B.1.1.7 (alpha), B.1.351 (beta), B.1.617.2 (delta), and B.1.1.529 (omicron). Our study describes a novel conserved epitope on the NTD that is readily targeted by vaccine-induced antibody responses. IMPORTANCE We report the first structure of a vaccine-induced antibody to SARS-CoV-2 spike isolated from plasmablasts 7 days after vaccination. The genetic sequence of the antibody PVI.V6-14 suggests that it is completely unmutated, meaning that this type of B cell did not undergo somatic hypermutation or affinity maturation; this cell was likely already present in the donor and was activated by the vaccine. This is, to our knowledge, also the first structure of an unmutated antibody in complex with its cognate antigen. PVI.V6-14 binds a novel, conserved epitope on the N-terminal domain (NTD) and neutralizes the original viral strain. PVI.V6-14 also binds the newly emerged variants B.1.1.7 (alpha), B.1.351 (beta), B.1.617.2 (delta), and B.1.1.529 (omicron). Given that this antibody was likely already present in the donor prior to vaccination, we believe that this antibody class could potentially "keep up" with the new variants, should they continue to emerge, by undergoing somatic hypermutation and affinity maturation.
Organizational Affiliation:
Department of Microbiology, Icahn School of Medicine at Mount Sinaigrid.59734.3c, New York, New York, USA.