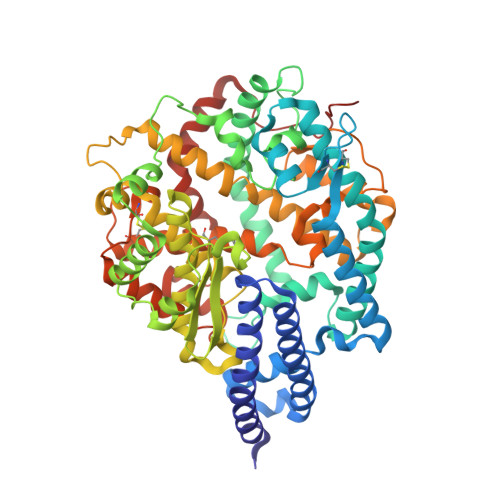
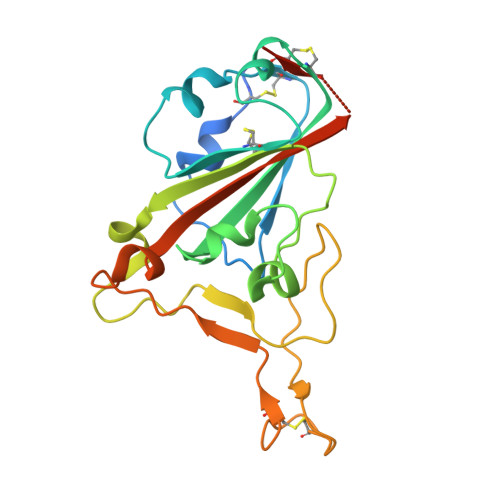
Structural Basis for Human Receptor Recognition by SARS-CoV-2 Omicron Variant BA.1.
Geng, Q., Shi, K., Ye, G., Zhang, W., Aihara, H., Li, F.(2022) J Virol 96: e0024922-e0024922
- PubMed: 35343765
- DOI: https://doi.org/10.1128/jvi.00249-22
- Primary Citation of Related Structures:
7U0N - PubMed Abstract:
The highly contagious and fast-spreading omicron variant of SARS-CoV-2 infects the respiratory tracts efficiently. The receptor-binding domain (RBD) of the omicron spike protein recognizes human angiotensin-converting enzyme 2 (ACE2) as its receptor and plays a critical role in the tissue tropism of SARS-CoV-2. Here, we showed that the omicron RBD (strain BA.1) binds to ACE2 more strongly than does the prototypic RBD from the original Wuhan strain. We also measured how individual omicron mutations affect ACE2 binding. We further determined the crystal structure of the omicron RBD (engineered to facilitate crystallization) complexed with ACE2 at 2.6 Å. The structure shows that omicron mutations caused significant structural rearrangements of two mutational hot spots at the RBD/ACE2 interface, elucidating how each omicron mutation affects ACE2 binding. The enhanced ACE2 binding by the omicron RBD may facilitate the omicron variant's infection of the respiratory tracts where ACE2 expression level is low. Our study provides insights into the receptor recognition and tissue tropism of the omicron variant. IMPORTANCE Despite the scarcity of the SARS-CoV-2 receptor-human angiotensin-converting enzyme 2 (ACE2)-in the respiratory tract, the omicron variant efficiently infects the respiratory tract, causing rapid and widespread infections of COVID-19. The omicron variant contains extensive mutations in the receptor-binding domain (RBD) of its spike protein that recognizes human ACE2. Here, using a combination of biochemical and X-ray crystallographic approaches, we showed that the omicron RBD binds to ACE2 with enhanced affinity and also elucidated the role of each of the omicron mutations in ACE2 binding. The enhanced ACE2 binding by the omicron RBD may contribute to the omicron variant's new viral tropism in the respiratory tract despite the low level of ACE2 expression in the tissue. These findings help us to understand tissue tropism of the omicron variant and shed light on the molecular evolution of SARS-CoV-2.
Organizational Affiliation:
Department of Veterinary and Biomedical Sciences, University of Minnesotagrid.17635.36, Saint Paul, Minnesota, USA.