Cryo-EM structure of SARS-CoV-2 S-Kappa variant (B.1.617.1), dimer of S trimer conformation 3
Yang, T.J., Yu, P.Y., Chang, Y.C., Hsu, S.T.D.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Spike glycoprotein | 1,283 | Severe acute respiratory syndrome coronavirus 2 | Mutation(s): 12 Gene Names: S, 2 | 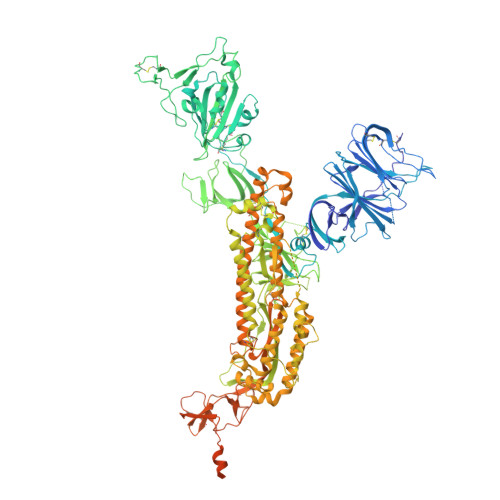 | |
UniProt | |||||
Find proteins for P0DTC2 (Severe acute respiratory syndrome coronavirus 2) Explore P0DTC2 Go to UniProtKB: P0DTC2 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0DTC2 | ||||
Glycosylation | |||||
| Glycosylation Sites: 16 | Go to GlyGen: P0DTC2-1 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | 3D Interactions |
| 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | AA [auth a], AB [auth 0], BA [auth b], BB [auth 1], CA [auth c], AA [auth a], AB [auth 0], BA [auth b], BB [auth 1], CA [auth c], CB [auth 2], DA [auth d], DB [auth 3], EA [auth e], EB [auth 4], FA [auth f], FB [auth 5], G, GA [auth g], H, HA [auth h], HB [auth 7], I, IB [auth 8], J, JA [auth j], JB [auth 9], KA [auth k], KB [auth AA], L, LA [auth l], LB [auth BA], M, MA [auth m], MB [auth CA], N, NA [auth n], NB [auth DA], O, OA [auth o], OB [auth EA], P, PA [auth p], PB [auth FA], Q, QA [auth q], QB [auth GA], R, RA [auth r], RB [auth HA], S, SA [auth s], T, TA [auth t], TB [auth JA], U, UB [auth KA], V, VA [auth v], VB [auth LA], WA [auth w], WB [auth MA], X, XA [auth x], XB [auth NA], Y, YA [auth y], YB [auth OA], Z, ZA [auth z], ZB [auth PA] | 2 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G42666HT GlyCosmos: G42666HT GlyGen: G42666HT | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | 3D Interactions |
| beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | GB [auth 6], IA [auth i], K, SB [auth IA], UA [auth u], GB [auth 6], IA [auth i], K, SB [auth IA], UA [auth u], W | 3 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G15407YE GlyCosmos: G15407YE GlyGen: G15407YE | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| NAG Query on NAG | AC [auth A] BC [auth A] CC [auth A] DC [auth A] EC [auth B] | 2-acetamido-2-deoxy-beta-D-glucopyranose C8 H15 N O6 OVRNDRQMDRJTHS-FMDGEEDCSA-N |  | ||
| Task | Software Package | Version |
|---|---|---|
| RECONSTRUCTION | cryoSPARC |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Ministry of Science and Technology (MoST, Taiwan) | Taiwan | 109-3114-Y-001-001 |
| Academia Sinica (Taiwan) | Taiwan | AS-CDA-109-L08 |
| Academia Sinica (Taiwan) | Taiwan | AS-CFII-108-110 |
| Academia Sinica (Taiwan) | Taiwan | AS-CFII108-111 |