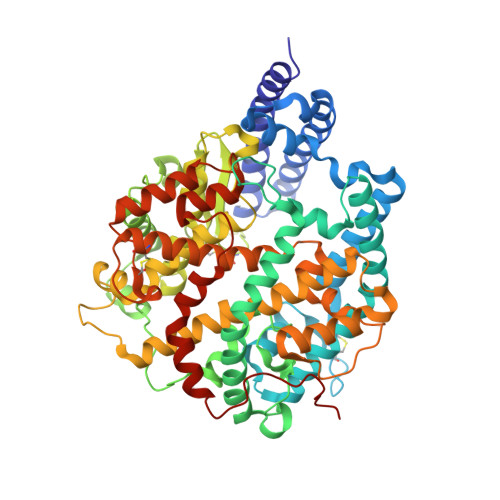
Broader-species receptor binding and structural bases of Omicron SARS-CoV-2 to both mouse and palm-civet ACE2s.
Li, L., Han, P., Huang, B., Xie, Y., Li, W., Zhang, D., Han, P., Xu, Z., Bai, B., Zhou, J., Kang, X., Li, X., Zheng, A., Zhang, R., Qiao, S., Zhao, X., Qi, J., Wang, Q., Liu, K., Gao, G.F.(2022) Cell Discov 8: 65-65
- PubMed: 35821014
- DOI: https://doi.org/10.1038/s41421-022-00431-0
- Primary Citation of Related Structures:
7WRH, 7WRI, 7WSK - PubMed Abstract:
The Omicron variant of SARS-CoV-2 carries multiple unusual mutations, particularly in the receptor-binding domain (RBD) of the spike (S) protein. Moreover, host-adapting mutations, such as residues 493, 498, and 501, were also observed in the Omicron RBD, which indicates that it is necessary to evaluate the interspecies transmission risk of the Omicron variant. Herein, we evaluated the interspecies recognition of the Omicron BA.1 and Delta RBDs by 27 ACE2 orthologs, including humans. We found that Omicron BA.1 expanded its receptor binding spectra to palm-civet, rodents, more bats (least horseshoe bat and greater horseshoe bat) and lesser hedgehog tenrec. Additionally, we determined the cryo-electron microscopy (cryo-EM) structure of the Omicron BA.1 S protein complexed with mouse ACE2 (mACE2) and the crystal structure of Omicron RBD complexed with palm-civet ACE2 (cvACE2). Several key residues for the host range have been identified. These results suggest that surveillance should be enhanced on the Omicron variant for its broader-species receptor binding to prevent spillover and expansion of reservoir hosts for a prolonged pandemic.
Organizational Affiliation:
CAS Key Laboratory of Pathogen Microbiology and Immunology, Institute of Microbiology, Chinese Academy of Sciences, Beijing, China.