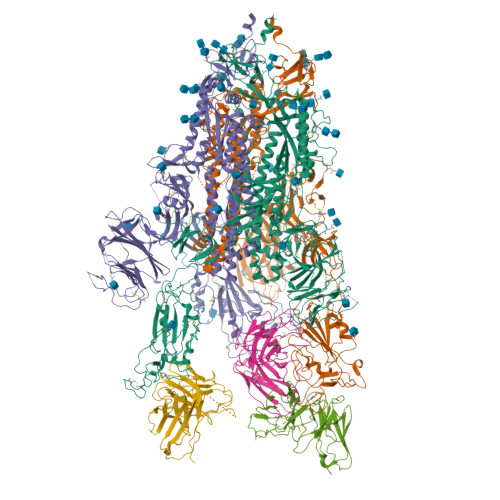
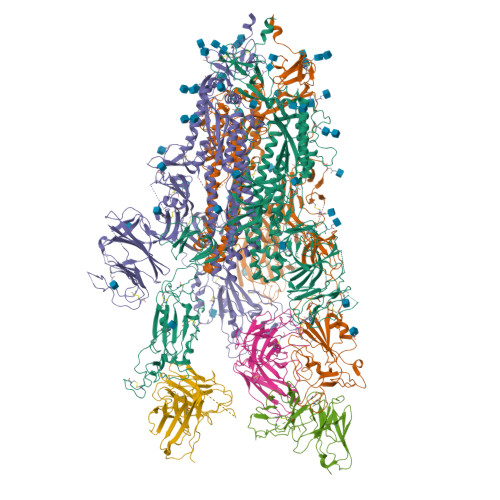
Spike mutation resilient scFv76 antibody counteracts SARS-CoV-2 lung damage upon aerosol delivery.
Milazzo, F.M., Chaves-Sanjuan, A., Minenkova, O., Santapaola, D., Anastasi, A.M., Battistuzzi, G., Chiapparino, C., Rosi, A., Merlo Pich, E., Albertoni, C., Marra, E., Luberto, L., Viollet, C., Spagnoli, L.G., Riccio, A., Rossi, A., Santoro, M.G., Ballabio, F., Paissoni, C., Camilloni, C., Bolognesi, M., De Santis, R.(2022) Mol Ther 31: 362-373
- PubMed: 36114671
- DOI: https://doi.org/10.1016/j.ymthe.2022.09.010
- Primary Citation of Related Structures:
7ZCE, 7ZCF - PubMed Abstract:
The uneven worldwide vaccination coverage against severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and emergence of variants escaping immunity call for broadly effective and easily deployable therapeutic agents. We have previously described the human single-chain scFv76 antibody, which recognizes SARS-CoV-2 Alpha, Beta, Gamma and Delta variants. We now show that scFv76 also neutralizes the infectivity and fusogenic activity of the Omicron BA.1 and BA.2 variants. Cryoelectron microscopy (cryo-EM) analysis reveals that scFv76 binds to a well-conserved SARS-CoV-2 spike epitope, providing the structural basis for its broad-spectrum activity. We demonstrate that nebulized scFv76 has therapeutic efficacy in a severe hACE2 transgenic mouse model of coronavirus disease 2019 (COVID-19) pneumonia, as shown by body weight and pulmonary viral load data. Counteraction of infection correlates with inhibition of lung inflammation, as observed by histopathology and expression of inflammatory cytokines and chemokines. Biomarkers of pulmonary endothelial damage were also significantly reduced in scFv76-treated mice. The results support use of nebulized scFv76 for COVID-19 induced by any SARS-CoV-2 variants that have emerged so far.
Organizational Affiliation:
Biotechnology R&D, Alfasigma SpA, Via Pontina Km 30.400, Pomezia, 00071 Rome, Italy.