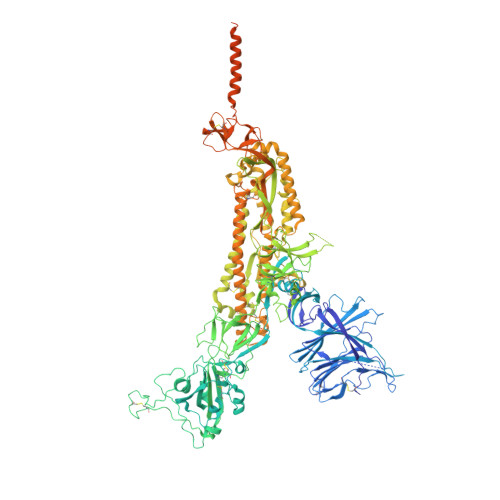
Structural and functional characteristics of the SARS-CoV-2 Omicron subvariant BA.2 spike protein.
Zhang, J., Tang, W., Gao, H., Lavine, C.L., Shi, W., Peng, H., Zhu, H., Anand, K., Kosikova, M., Kwon, H.J., Tong, P., Gautam, A., Rits-Volloch, S., Wang, S., Mayer, M.L., Wesemann, D.R., Seaman, M.S., Lu, J., Xiao, T., Xie, H., Chen, B.(2023) Nat Struct Mol Biol 30: 980-990
- PubMed: 37430064
- DOI: https://doi.org/10.1038/s41594-023-01023-6
- Primary Citation of Related Structures:
8D55, 8D56, 8D5A - PubMed Abstract:
The Omicron subvariant BA.2 has become the dominant circulating strain of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) in many countries. Here, we have characterized structural, functional and antigenic properties of the full-length BA.2 spike (S) protein and compared replication of the authentic virus in cell culture and an animal model with previously prevalent variants. BA.2 S can fuse membranes slightly more efficiently than Omicron BA.1, but still less efficiently than other previous variants. Both BA.1 and BA.2 viruses replicated substantially faster in animal lungs than the early G614 (B.1) strain in the absence of pre-existing immunity, possibly explaining the increased transmissibility despite their functionally compromised spikes. As in BA.1, mutations in the BA.2 S remodel its antigenic surfaces, leading to strong resistance to neutralizing antibodies. These results suggest that both immune evasion and replicative advantage may contribute to the heightened transmissibility of the Omicron subvariants.
Organizational Affiliation:
Division of Molecular Medicine, Boston Children's Hospital, Boston, MA, USA.