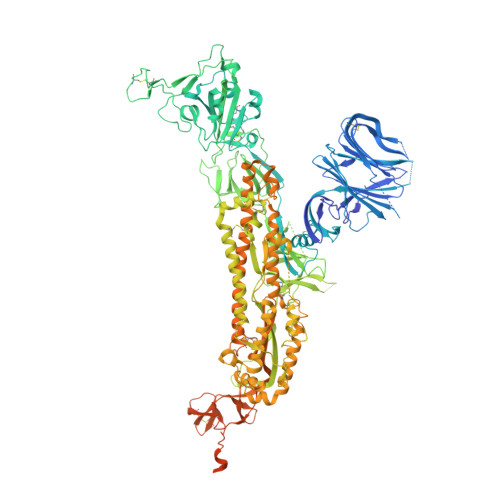
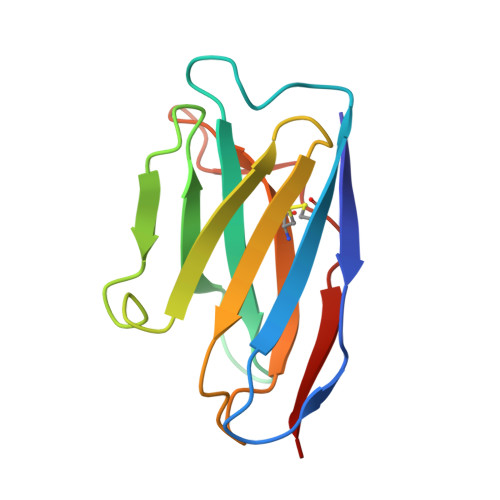
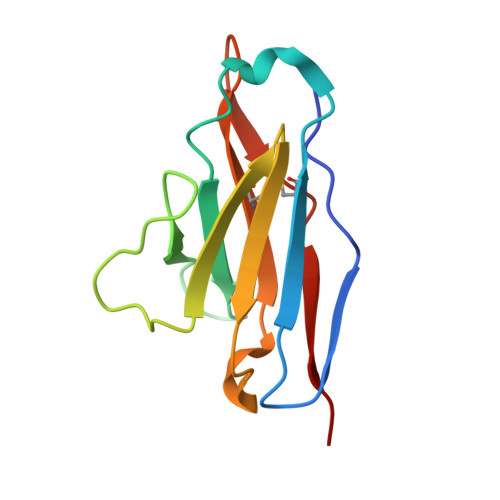
Imprinted antibody responses against SARS-CoV-2 Omicron sublineages.
Park, Y.J., Pinto, D., Walls, A.C., Liu, Z., De Marco, A., Benigni, F., Zatta, F., Silacci-Fregni, C., Bassi, J., Sprouse, K.R., Addetia, A., Bowen, J.E., Stewart, C., Giurdanella, M., Saliba, C., Guarino, B., Schmid, M.A., Franko, N.M., Logue, J.K., Dang, H.V., Hauser, K., di Iulio, J., Rivera, W., Schnell, G., Rajesh, A., Zhou, J., Farhat, N., Kaiser, H., Montiel-Ruiz, M., Noack, J., Lempp, F.A., Janer, J., Abdelnabi, R., Maes, P., Ferrari, P., Ceschi, A., Giannini, O., de Melo, G.D., Kergoat, L., Bourhy, H., Neyts, J., Soriaga, L., Purcell, L.A., Snell, G., Whelan, S.P.J., Lanzavecchia, A., Virgin, H.W., Piccoli, L., Chu, H.Y., Pizzuto, M.S., Corti, D., Veesler, D.(2022) Science 378: 619-627
- PubMed: 36264829
- DOI: https://doi.org/10.1126/science.adc9127
- Primary Citation of Related Structures:
8ERQ, 8ERR - PubMed Abstract:
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) Omicron sublineages carry distinct spike mutations resulting in escape from antibodies induced by previous infection or vaccination. We show that hybrid immunity or vaccine boosters elicit plasma-neutralizing antibodies against Omicron BA.1, BA.2, BA.2.12.1, and BA.4/5, and that breakthrough infections, but not vaccination alone, induce neutralizing antibodies in the nasal mucosa. Consistent with immunological imprinting, most antibodies derived from memory B cells or plasma cells of Omicron breakthrough cases cross-react with the Wuhan-Hu-1, BA.1, BA.2, and BA.4/5 receptor-binding domains, whereas Omicron primary infections elicit B cells of narrow specificity up to 6 months after infection. Although most clinical antibodies have reduced neutralization of Omicron, we identified an ultrapotent pan-variant-neutralizing antibody that is a strong candidate for clinical development.
Organizational Affiliation:
Department of Biochemistry, University of Washington, Seattle, WA, USA.