Distinct in vitro and in vivo neutralization profiles of monoclonal antibodies elicited by the receptor binding domain of the ancestral SARS-CoV-2.
Kwon, H.J., Zhang, J., Kosikova, M., Tang, W., Ortega-Rodriguez, U., Peng, H., Meseda, C.A., Pedro, C.L., Schmeisser, F., Lu, J., Kang, I., Zhou, B., Davis, C.T., Wentworth, D.E., Chen, W.H., Shriver, M.C., Barnes, R.S., Pasetti, M.F., Weir, J.P., Chen, B., Xie, H.(2023) J Med Virol 95: e28673-e28673
- PubMed: 36916782
- DOI: https://doi.org/10.1002/jmv.28673
- Primary Citation of Related Structures:
8GJM, 8GJN - PubMed Abstract:
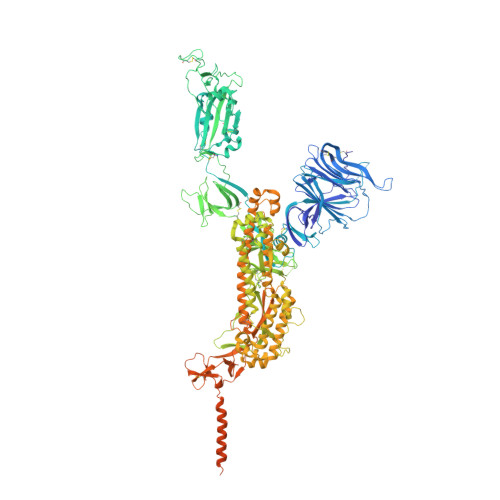
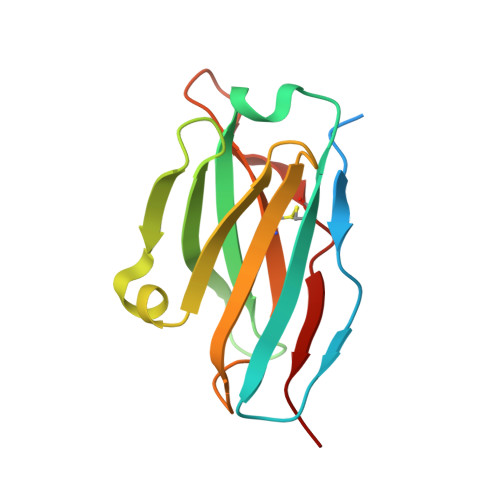
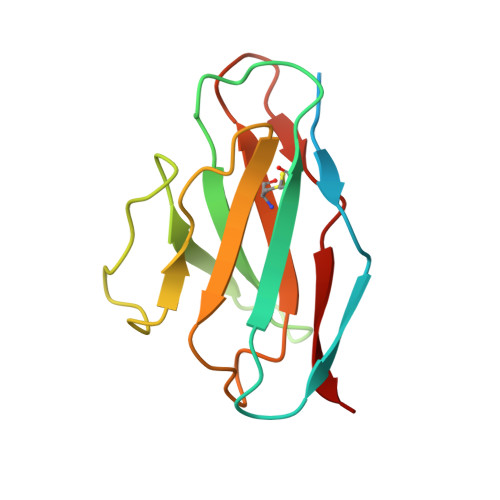
Broadly neutralizing antibodies against severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) variants are sought to curb coronavirus disease 2019 (COVID-19) infections. Here we produced and characterized a set of mouse monoclonal antibodies (mAbs) specific for the ancestral SARS-CoV-2 receptor binding domain (RBD). Two of them, 17A7 and 17B10, were highly potent in microneutralization assay with 50% inhibitory concentration (IC 50 ) ≤135 ng/mL against infectious SARS-CoV-2 variants, including G614, Alpha, Beta, Gamma, Delta, Epsilon, Zeta, Kappa, Lambda, B.1.1.298, B.1.222, B.1.5, and R.1. Both mAbs (especially 17A7) also exhibited strong in vivo efficacy in protecting K18-hACE2 transgenic mice from the lethal infection with G614, Alpha, Beta, Gamma, and Delta viruses. Structural analysis indicated that 17A7 and 17B10 target the tip of the receptor binding motif in the RBD-up conformation. A third RBD-reactive mAb (3A6) although escaped by Beta and Gamma, was highly effective in cross-neutralizing Delta and Omicron BA.1 variants in vitro and in vivo. In competition experiments, antibodies targeting epitopes similar to these 3 mAbs were rarely enriched in human COVID-19 convalescent sera or postvaccination sera. These results are helpful to inform new antibody/vaccine design and these mAbs can be useful tools for characterizing SARS-CoV-2 variants and elicited antibody responses.
Organizational Affiliation:
Laboratory of Pediatric and Respiratory Viral Diseases, Division of Viral Products, Center for Biologics Evaluation and Research, Office of Vaccines Research and Review, United States Food and Drug Administration, Silver Spring, Maryland, USA.