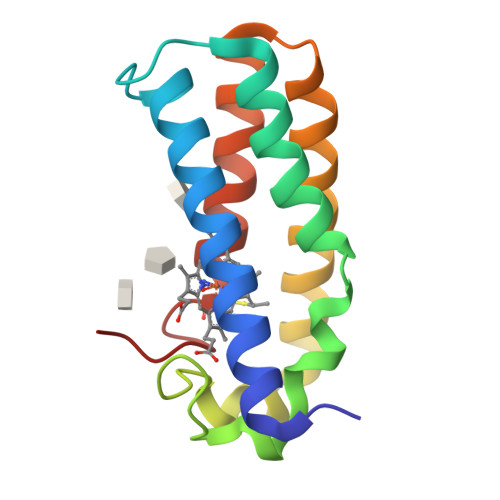
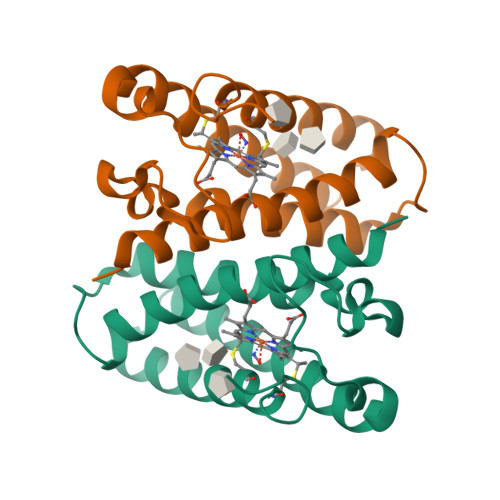
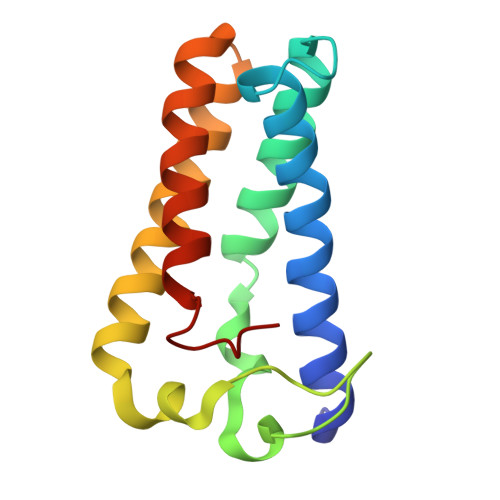
Conformational rigidity of cytochrome c'-alpha from a thermophile is associated with slow NO binding.
Fujii, S., Wilson, M.T., Adams, H.R., Mikolajek, H., Svistunenko, D.A., Smyth, P., Andrew, C.R., Sambongi, Y., Hough, M.A.(2024) Biophys J 123: 2594-2603
- PubMed: 38937973
- DOI: https://doi.org/10.1016/j.bpj.2024.06.026
- Primary Citation of Related Structures:
8RKP - PubMed Abstract:
Cytochromes c'-α are nitric oxide (NO)-binding heme proteins derived from bacteria that can thrive in a wide range of temperature environments. Studies of mesophilic Alcaligenes xylosoxidans cytochrome c'-α (AxCP-α) have revealed an unusual NO-binding mechanism involving both heme faces, in which NO first binds to form a distal hexa-coordinate Fe(II)-NO (6cNO) intermediate and then displaces the proximal His to form a proximal penta-coordinate Fe(II)-NO (5cNO) final product. Here, we characterize a thermally stable cytochrome c'-α from thermophilic Hydrogenophilus thermoluteolus (PhCP-α) to understand how protein thermal stability affects NO binding. Electron paramagnetic and resonance Raman spectroscopies reveal the formation of a PhCP-α 5cNO product, with time-resolved (stopped-flow) UV-vis absorbance indicating the involvement of a 6cNO intermediate. Relative to AxCP-α, the rates of 6cNO and 5cNO formation in PhCP-α are ∼11- and ∼13-fold lower, respectively. Notably, x-ray crystal structures of PhCP-α in the presence and absence of NO suggest that the sluggish formation of the proximal 5cNO product results from conformational rigidity: the Arg-132 residue (adjacent to the proximal His ligand) is held in place by a salt bridge between Arg-75 and Glu-135 (an interaction not present in AxCP-α or a psychrophilic counterpart). Overall, our data provide fresh insights into structural factors controlling NO binding in heme proteins, including 5cNO complexes relevant to eukaryotic NO sensors.
Organizational Affiliation:
Diamond Light Source Ltd, Harwell Science and Innovation Campus, Didcot, United Kingdom; Research Complex at Harwell, Harwell Science and Innovation Campus, Didcot, United Kingdom; Graduate School of Integrated Sciences for Life, Hiroshima University, Higashi-Hiroshima, Japan. Electronic address: sofuji@hiroshima-u.ac.jp.