Bifunctional Inhibition of Neutrophil Elastase and Cathepsin-G by Eap4 from S. aureus
Mishra, N.B., Geisbrecht, B.V.To be published.
Experimental Data Snapshot
Starting Models: experimental
View more details
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Neutrophil elastase | A, B [auth D], F [auth G], J | 218 | Homo sapiens | Mutation(s): 0 EC: 3.4.21.37 | 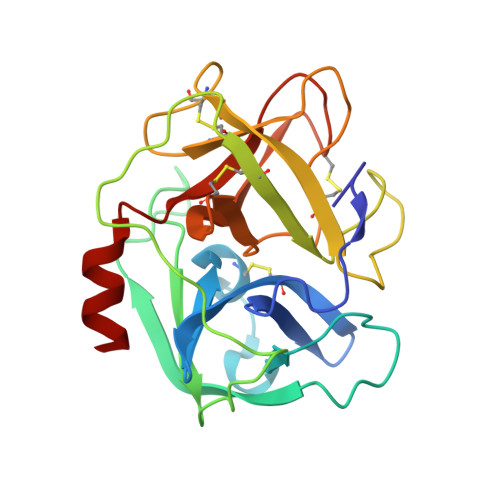 |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P08246 (Homo sapiens) Explore P08246 Go to UniProtKB: P08246 | |||||
PHAROS: P08246 GTEx: ENSG00000197561 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P08246 | ||||
Glycosylation | |||||
| Glycosylation Sites: 2 | Go to GlyGen: P08246-1 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Extracellular Adherence Protein | C, E [auth F], H [auth I], L | 108 | Staphylococcus aureus subsp. aureus Mu50 | Mutation(s): 0 Gene Names: map, SAV1938 | 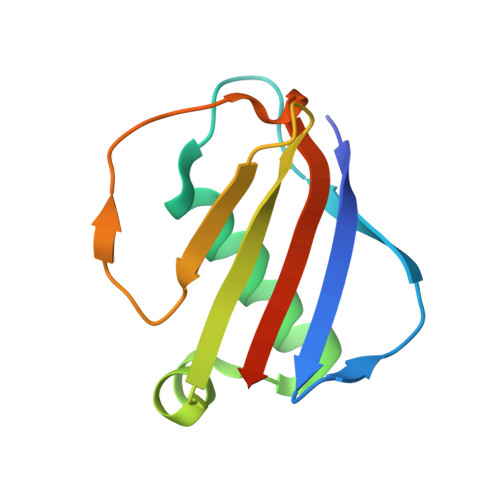 |
UniProt | |||||
Find proteins for Q99QS1 (Staphylococcus aureus (strain Mu50 / ATCC 700699)) Explore Q99QS1 Go to UniProtKB: Q99QS1 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q99QS1 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Cathepsin-G | D [auth E], G [auth H], I [auth B], K | 223 | Homo sapiens | Mutation(s): 0 | 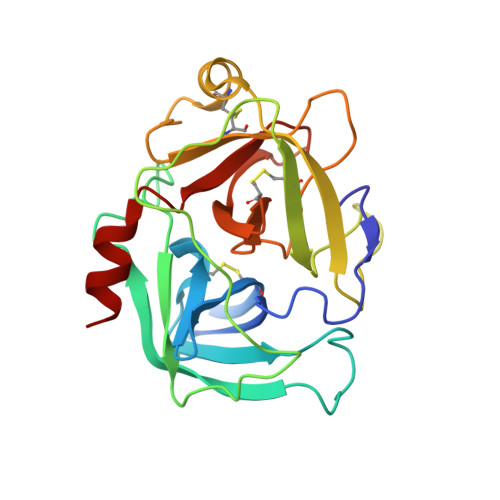 |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P08311 (Homo sapiens) Explore P08311 Go to UniProtKB: P08311 | |||||
PHAROS: P08311 GTEx: ENSG00000100448 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P08311 | ||||
Glycosylation | |||||
| Glycosylation Sites: 1 | Go to GlyGen: P08311-1 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | 3D Interactions |
| beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose | M, O, P, Q | 4 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G32152BH GlyCosmos: G32152BH GlyGen: G32152BH | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| NAG Query on NAG | U [auth E], V [auth H], W [auth B], X [auth K] | 2-acetamido-2-deoxy-beta-D-glucopyranose C8 H15 N O6 OVRNDRQMDRJTHS-FMDGEEDCSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 82.207 | α = 82.77 |
| b = 85.527 | β = 85.23 |
| c = 89.536 | γ = 83.6 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| HKL-2000 | data scaling |
| HKL-2000 | data reduction |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Institute of General Medical Sciences (NIH/NIGMS) | United States | R35GM140852 |