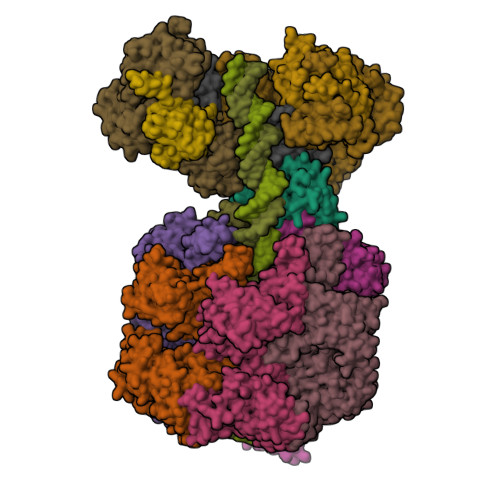
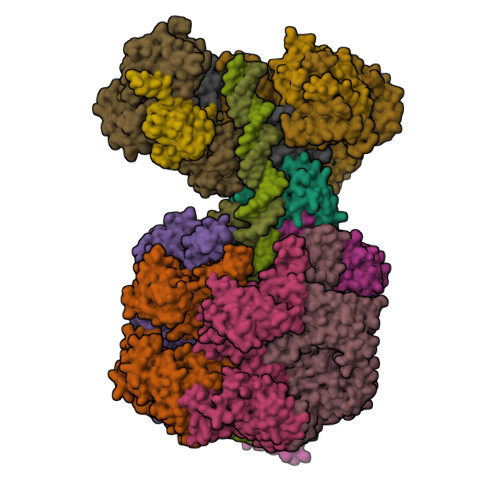
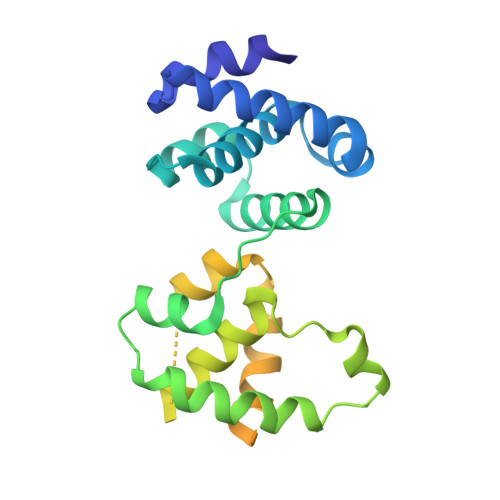
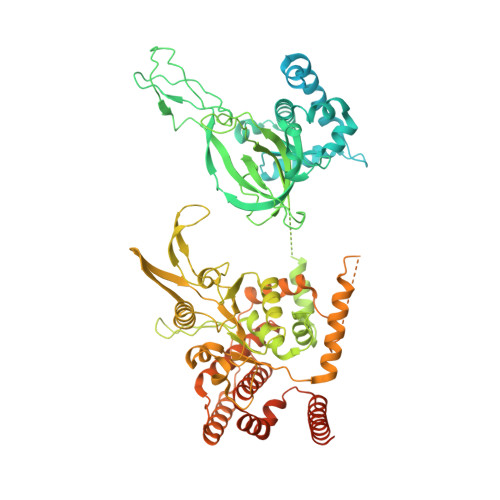
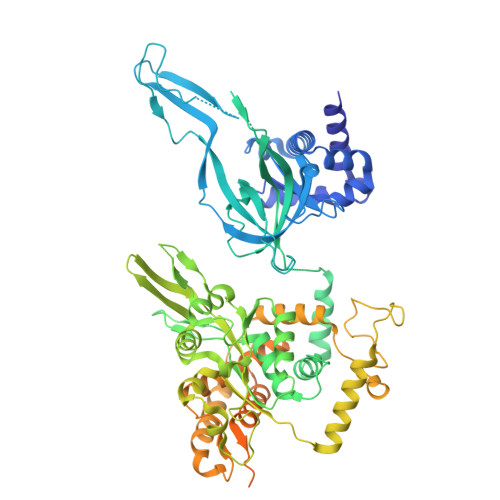
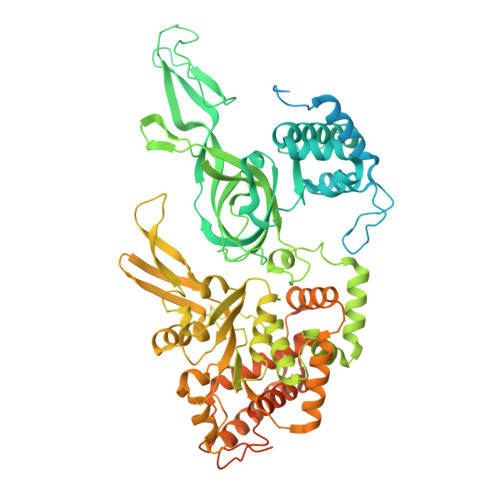
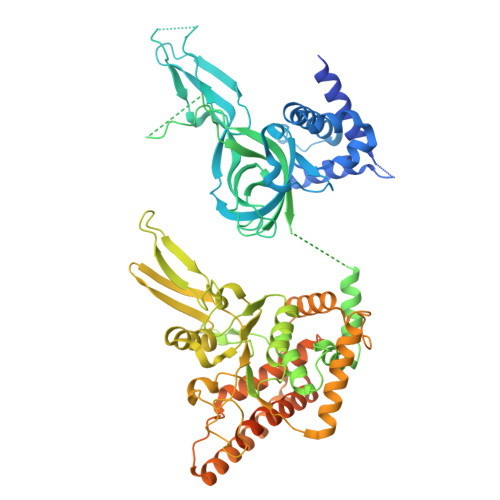
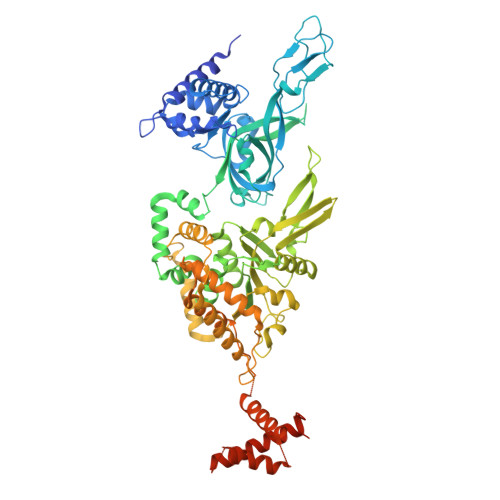
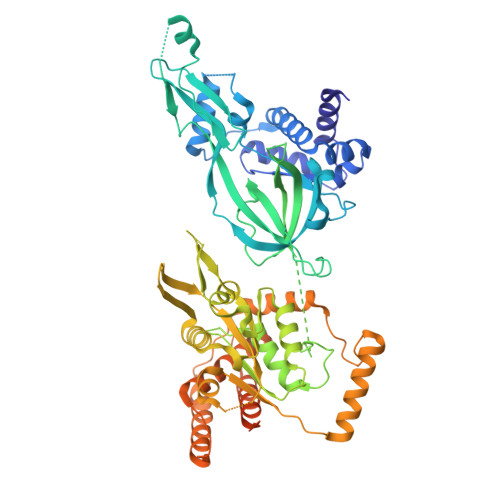
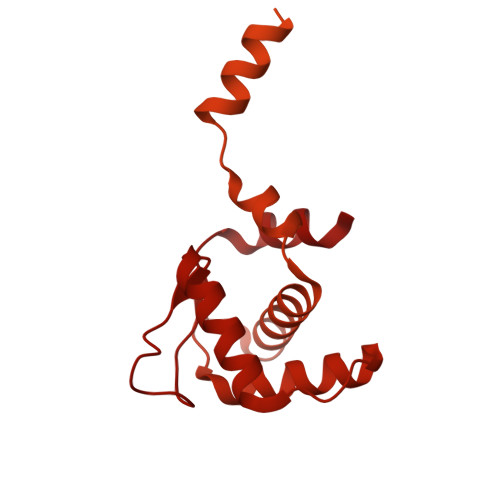
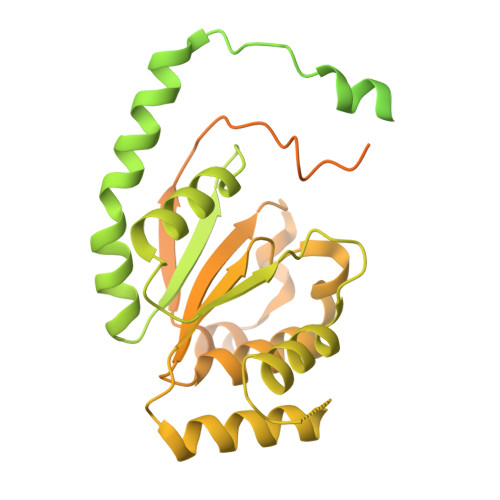
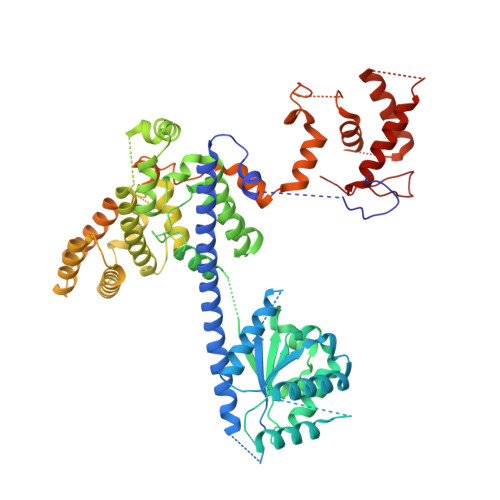
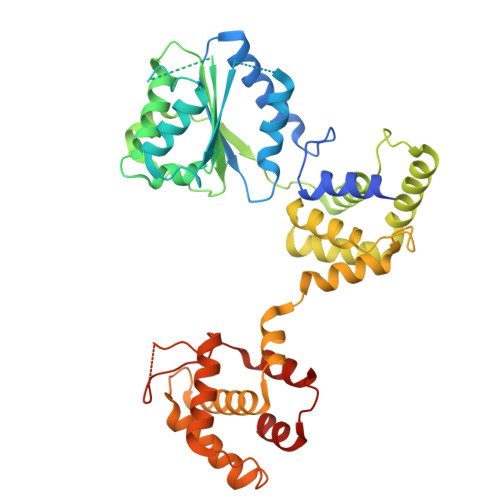
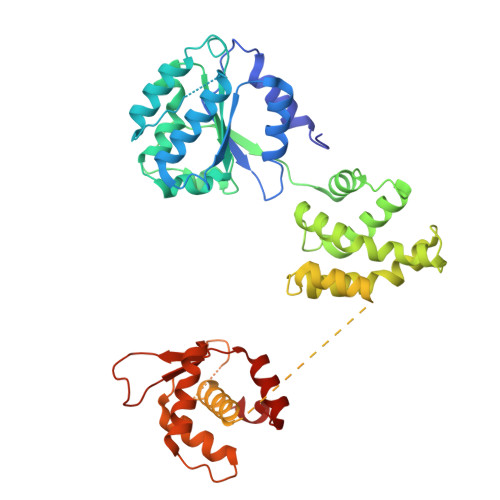
MCM double hexamer loading visualized with human proteins.
Weissmann, F., Greiwe, J.F., Puhringer, T., Eastwood, E.L., Couves, E.C., Miller, T.C.R., Diffley, J.F.X., Costa, A.(2024) Nature 636: 499-508
- PubMed: 39604733
- DOI: https://doi.org/10.1038/s41586-024-08263-6
- Primary Citation of Related Structures:
8S09, 8S0A, 8S0B, 8S0C, 8S0D, 8S0E, 8S0F - PubMed Abstract:
Eukaryotic DNA replication begins with the loading of the MCM replicative DNA helicase as a head-to-head double hexamer at origins of DNA replication 1-3 . Our current understanding of how the double hexamer is assembled by the origin recognition complex (ORC), CDC6 and CDT1 comes mostly from budding yeast. Here we characterize human double hexamer (hDH) loading using biochemical reconstitution and cryo-electron microscopy with purified proteins. We show that the human double hexamer engages DNA differently from the yeast double hexamer (yDH), and generates approximately five base pairs of underwound DNA at the interface between hexamers, as seen in hDH isolated from cells 4 . We identify several differences from the yeast double hexamer in the order of factor recruitment and dependencies during hDH assembly. Unlike in yeast 5-8 , the ORC6 subunit of the ORC is not essential for initial MCM recruitment or hDH loading, but contributes to an alternative hDH assembly pathway that requires an intrinsically disordered region in ORC1, which may work through a MCM-ORC intermediate. Our work presents a detailed view of how double hexamers are assembled in an organism that uses sequence-independent replication origins, provides further evidence for diversity in eukaryotic double hexamer assembly mechanisms 9 , and represents a first step towards reconstitution of DNA replication initiation with purified human proteins.
Organizational Affiliation:
Chromosome Replication Laboratory, The Francis Crick Institute, London, UK.