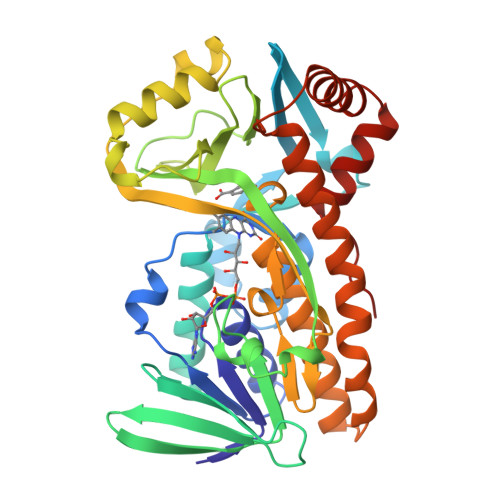
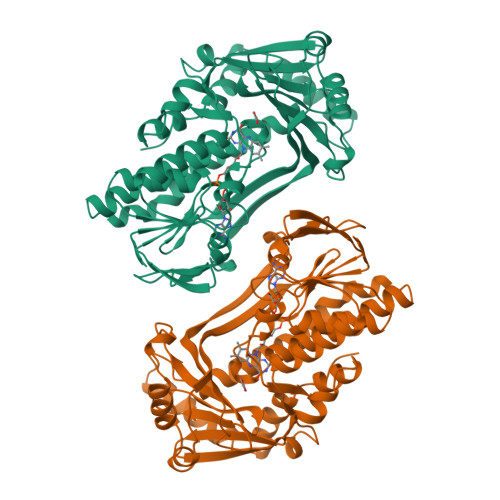
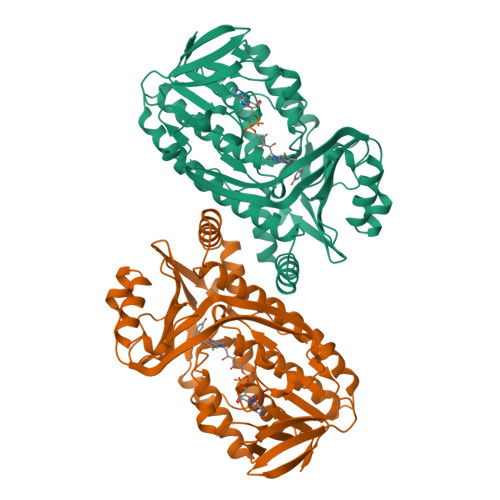
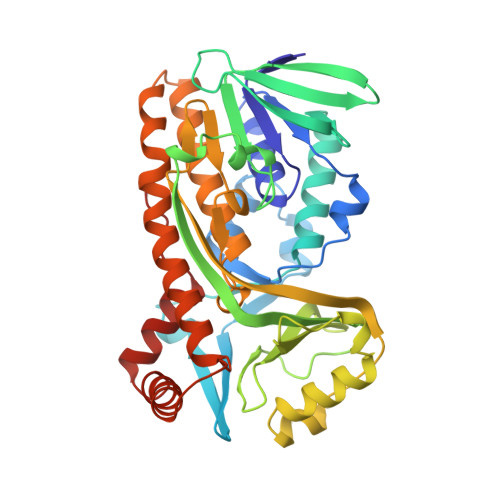
Switch of coenzyme specificity of p-hydroxybenzoate hydroxylase.
Eppink, M.H., Overkamp, K.M., Schreuder, H.A., Van Berkel, W.J.(1999) J Mol Biol 292: 87-96
- PubMed: 10493859
- DOI: https://doi.org/10.1006/jmbi.1999.3015
- Primary Citation of Related Structures:
1CJ2, 1CJ3, 1CJ4 - PubMed Abstract:
p-Hydroxybenzoate hydroxylase (PHBH) is the archetype of the family of NAD(P)H-dependent flavoprotein aromatic hydroxylases. These enzymes share a conserved FAD-binding domain but lack a recognizable fold for binding the pyridine nucleotide. We have switched the coenzyme specificity of strictly NADPH-dependent PHBH from Pseudomonas fluorescens by site-directed mutagenesis. To that end, we altered the solvent exposed helix H2 region (residues 33-40) of the FAD-binding domain. Non-conservative selective replacements of Arg33 and Tyr38 weakened the binding of NADPH without disturbing the protein architecture. Introduction of a basic residue at position 34 increased the NADPH binding strength. Double (M2) and quadruple (M4) substitutions in the N-terminal part of helix H2 did not change the coenzyme specificity. By extending the replacements towards residues 38 and 40, M5 and M6 mutants were generated which were catalytically more efficient with NADH than with NADPH. It is concluded that specificity in P. fluorescens PHBH is conferred by interactions of Arg33, Tyr38 and Arg42 with the 2'-phosphate moiety of bound NADPH, and that introduction of an acidic group at position 38 potentially enables the recognition of the 2'-hydroxy group of NADH. This is the first report on the coenzyme reversion of a flavoprotein aromatic hydroxylase.
Organizational Affiliation:
Department of Biomolecular Sciences, Laboratory of Biochemistry, Wageningen University, Wageningen, 6703 HA, The Netherlands.