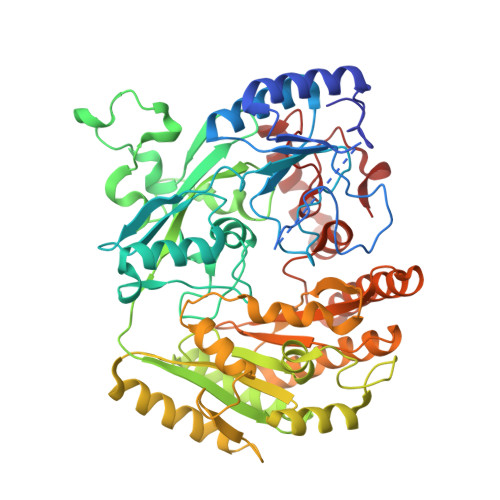
Structural Analysis of Flavinylation in Vanillyl-Alcohol Oxidase
Fraaije, M.W., Van Der Heuvel, R.H.H., Van Berkel, W.J.H., Mattevi, A.(2001) J Biol Chem 275: 38654
- PubMed: 10984479
- DOI: https://doi.org/10.1074/jbc.M004753200
- Primary Citation of Related Structures:
1E8F, 1E8G, 1E8H - PubMed Abstract:
Vanillyl-alcohol oxidase (VAO) is member of a newly recognized flavoprotein family of structurally related oxidoreductases. The enzyme contains a covalently linked FAD cofactor. To study the mechanism of flavinylation we have created a design point mutation (His-61 --> Thr). In the mutant enzyme the covalent His-C8alpha-flavin linkage is not formed, while the enzyme is still able to bind FAD and perform catalysis. The H61T mutant displays a similar affinity for FAD and ADP (K(d) = 1.8 and 2.1 microm, respectively) but does not interact with FMN. H61T is about 10-fold less active with 4-(methoxymethyl)phenol) (k(cat) = 0.24 s(-)(1), K(m) = 40 microm) than the wild-type enzyme. The crystal structures of both the holo and apo form of H61T are highly similar to the structure of wild-type VAO, indicating that binding of FAD to the apoprotein does not require major structural rearrangements. These results show that covalent flavinylation is an autocatalytical process in which His-61 plays a crucial role by activating His-422. Furthermore, our studies clearly demonstrate that in VAO, the FAD binds via a typical lock-and-key approach to a preorganized binding site.
Organizational Affiliation:
Department of Biomolecular Sciences, Laboratory of Biochemistry, Wageningen University, Dreijenlaan 3, 6703 HA Wageningen, The Netherlands. m.w.fraaije@chem.rug.nl