The Active Site of Cellobiohydrolase Cel6A from Trichoderma Reesei: The Roles of Aspartic Acids D221 and D175
Koivula, A., Ruohonen, L., Wohlfahrt, G., Reinikainen, T., Teeri, T.T., Piens, K., Claeyssens, M., Weber, M., Vasella, A., Becker, D., Sinnott, M.L., Zou, J.-Y., Kleywegt, G.J., Szardenings, M., Stahlberg, J., Jones, T.A.(2002) J Am Chem Soc 124: 10015
- PubMed: 12188666
- DOI: https://doi.org/10.1021/ja012659q
- Primary Citation of Related Structures:
1HGW, 1HGY - PubMed Abstract:
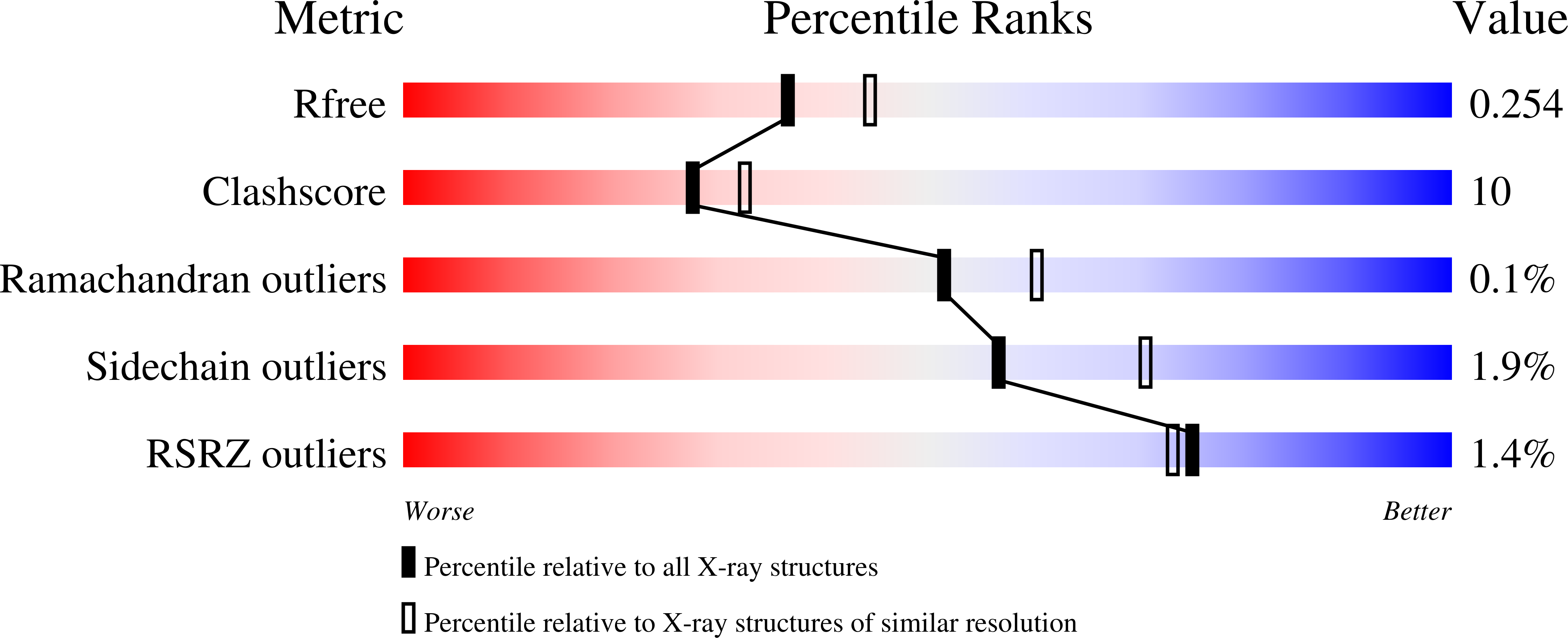
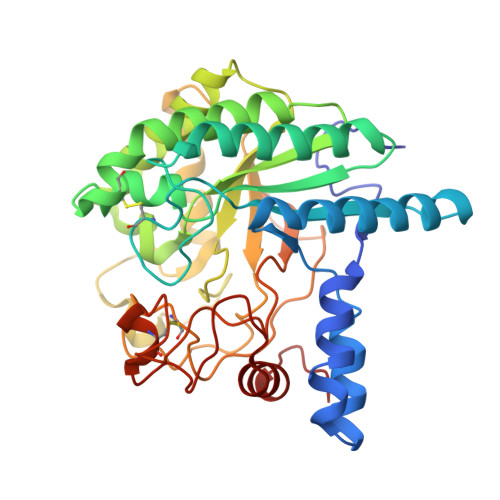
Trichoderma reesei cellobiohydrolase Cel6A is an inverting glycosidase. Structural studies have established that the tunnel-shaped active site of Cel6A contains two aspartic acids, D221 and D175, that are close to the glycosidic oxygen of the scissile bond and at hydrogen-bonding distance from each other. Here, site-directed mutagenesis, X-ray crystallography, and enzyme kinetic studies have been used to confirm the role of residue D221 as the catalytic acid. D175 is shown to affect protonation of D221 and to contribute to the electrostatic stabilization of the partial positive charge in the transition state. Structural and modeling studies suggest that the single-displacement mechanism of Cel6A may not directly involve a catalytic base. The value of (D2O)(V) of 1.16 +/- 0.14 for hydrolysis of cellotriose suggests that the large direct effect expected for proton transfer from the nucleophilic water through a water chain (Grotthus mechanism) is offset by an inverse effect arising from reversibly breaking the short, tight hydrogen bond between D221 and D175 before catalysis.
Organizational Affiliation:
VTT Biotechnology, P.O. Box 1500, FIN-02044 VTT, Espoo, Finland.