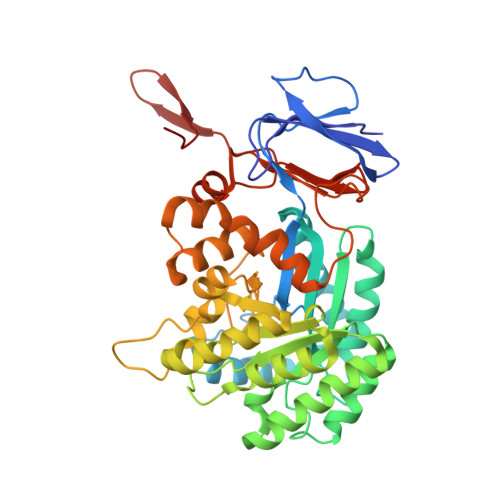
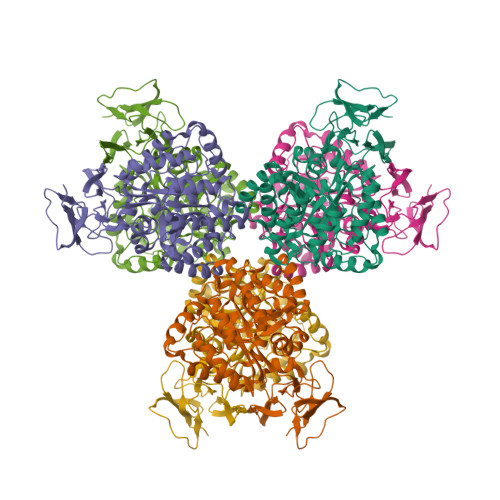
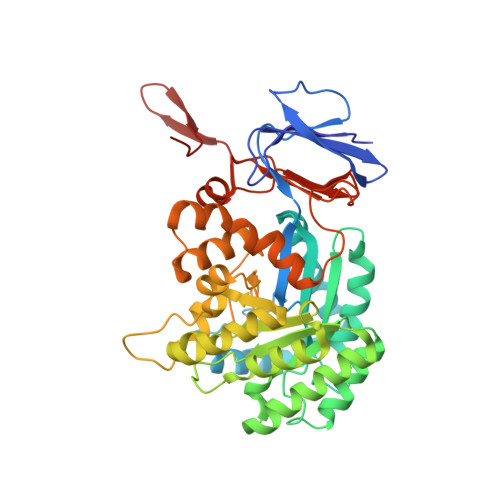
The structure of Escherichia coli cytosine deaminase.
Ireton, G.C., McDermott, G., Black, M.E., Stoddard, B.L.(2002) J Mol Biology 315: 687-697
- PubMed: 11812140
- DOI: https://doi.org/10.1006/jmbi.2001.5277
- Primary Citation of Related Structures:
1K6W, 1K70 - PubMed Abstract:
Cytosine deaminase (CD) catalyzes the deamination of cytosine, producing uracil. This enzyme is present in prokaryotes and fungi (but not multicellular eukaryotes) and is an important member of the pyrimidine salvage pathway in those organisms. The same enzyme also catalyzes the conversion of 5-fluorocytosine to 5-fluorouracil; this activity allows the formation of a cytotoxic chemotherapeutic agent from a non-cytotoxic precursor. The enzyme is of widespread interest both for antimicrobial drug design and for gene therapy applications against tumors. The structure of Escherichia coli CD has been determined in the presence and absence of a bound mechanism-based inhibitor. The enzyme forms an (alphabeta)(8) barrel structure with structural similarity to adenosine deaminase, a relationship that is undetectable at the sequence level, and no similarity to bacterial cytidine deaminase. The enzyme is packed into a hexameric assembly stabilized by a unique domain-swapping interaction between enzyme subunits. The active site is located in the mouth of the enzyme barrel and contains a bound iron ion that coordinates a hydroxyl nucleophile. Substrate binding involves a significant conformational change that sequesters the reaction complex from solvent.
Organizational Affiliation:
Fred Hutchinson Cancer Research Center and the Graduate Program in Molecular and Cell Biology, University of Washington, 1100 Fairview Ave. N. A3-023, Seattle, WA 98109, USA.