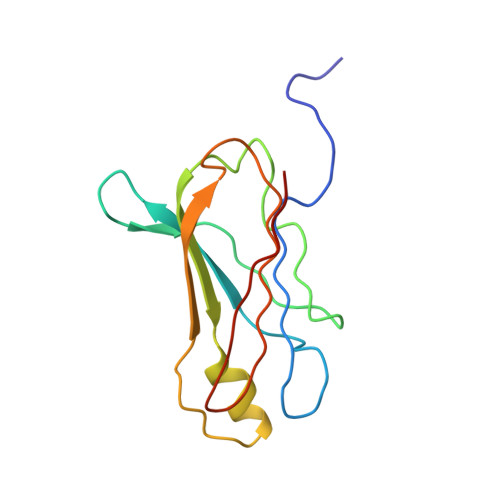
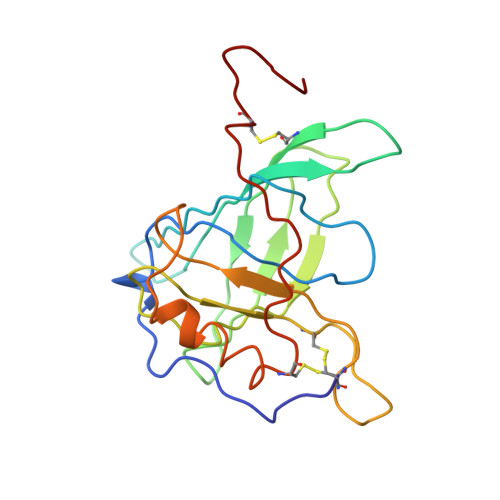
The structure of human retinol-binding protein (RBP) with its carrier protein transthyretin reveals an interaction with the carboxy terminus of RBP.
Naylor, H.M., Newcomer, M.E.(1999) Biochemistry 38: 2647-2653
- PubMed: 10052934
- DOI: https://doi.org/10.1021/bi982291i
- Primary Citation of Related Structures:
1QAB - PubMed Abstract:
Whether ultimately utilized as retinoic acid, retinal, or retinol, vitamin A is transported to the target cells as all-trans-retinol bound to retinol-binding protein (RBP). Circulating in the plasma, RBP itself is bound to transthyretin (TTR, previously referred to as thyroxine-binding prealbumin). In vitro one tetramer of TTR can bind two molecules of retinol-binding protein. However, the concentration of RBP in the plasma is limiting, and the complex isolated from serum is composed of TTR and RBP in a 1 to 1 stoichiometry. We report here the crystallographic structure at 3.2 A of the protein-protein complex of human RBP and TTR. RBP binds at a 2-fold axis of symmetry in the TTR tetramer, and consequently the recognition site itself has 2-fold symmetry: Four TTR amino acids (Arg-21, Val-20, Leu-82, and Ile-84) are contributed by two monomers. Amino acids Trp-67, Phe-96, and Leu-63 and -97 from RBP are flanked by the symmetry-related side chains from TTR. In addition, the structure reveals an interaction of the carboxy terminus of RBP at the protein-protein recognition interface. This interaction, which involves Leu-182 and Leu-183 of RBP, is consistent with the observation that naturally occurring truncated forms of the protein are more readily cleared from plasma than full-length RBP. Complex formation prevents extensive loss of RBP through glomerular filtration, and the loss of Leu-182 and Leu-183 would result in a decreased affinity of RBP for TTR.
Organizational Affiliation:
Department of Biochemistry, Vanderbilt University School of Medicine, Nashville, Tennessee 37232-0146, USA.