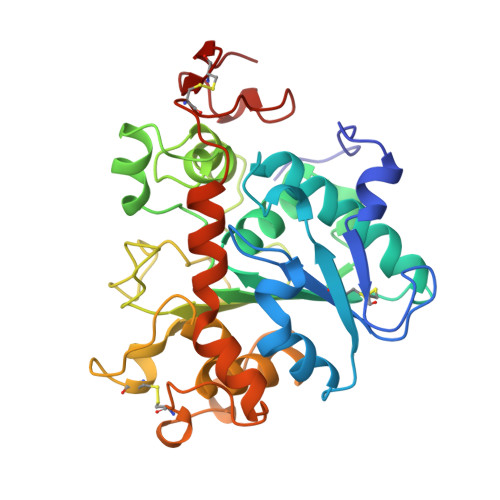
The sequence, crystal structure determination and refinement of two crystal forms of lipase B from Candida antarctica.
Uppenberg, J., Hansen, M.T., Patkar, S., Jones, T.A.(1994) Structure 2: 293-308
- PubMed: 8087556
- DOI: https://doi.org/10.1016/s0969-2126(00)00031-9
- Primary Citation of Related Structures:
1TCA, 1TCB, 1TCC - PubMed Abstract:
Lipases constitute a family of enzymes that hydrolyze triglycerides. They occur in many organisms and display a wide variety of substrate specificities. In recent years, much progress has been made towards explaining the mechanism of these enzymes and their ability to hydrolyze their substrates at an oil-water interface. We have determined the DNA and amino acid sequences for lipase B from the yeast Candida antarctica. The primary sequence has no significant homology to any other known lipase and deviates from the consensus sequence around the active site serine that is found in other lipases. We have determined the crystal structure of this enzyme using multiple isomorphous replacement methods for two crystal forms. Models for the orthorhombic and monoclinic crystal forms of the enzyme have been refined to 1.55 A and 2.1 A resolution, respectively. Lipase B is an alpha/beta type protein that has many features in common with previously determined lipase structures and other related enzymes. In the monoclinic crystal form, lipid-like molecules, most likely beta-octyl glucoside, can be seen close to the active site. The behaviour of these lipid molecules in the crystal structure has been studied at different pH values. The structure of Candida antarctica lipase B shows that the enzyme has a Ser-His-Asp catalytic triad in its active site. The structure appears to be in an 'open' conformation with a rather restricted entrance to the active site. We believe that this accounts for the substrate specificity and high degree of stereospecificity of this lipase.
Organizational Affiliation:
Department of Molecular Biology, Uppsala University, Sweden.