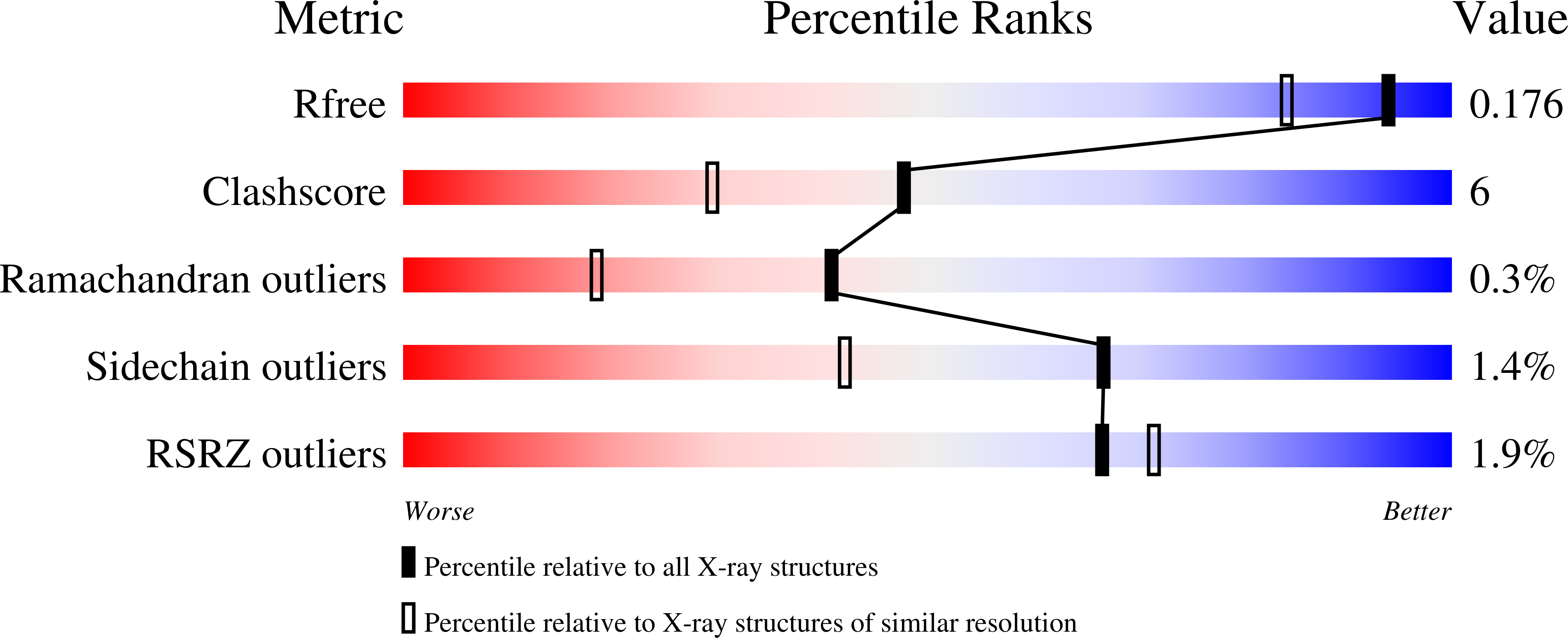
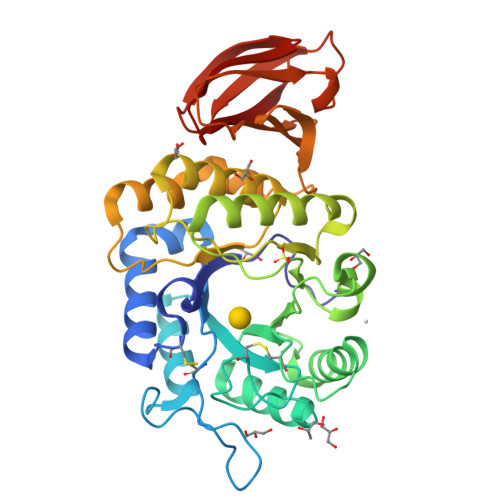
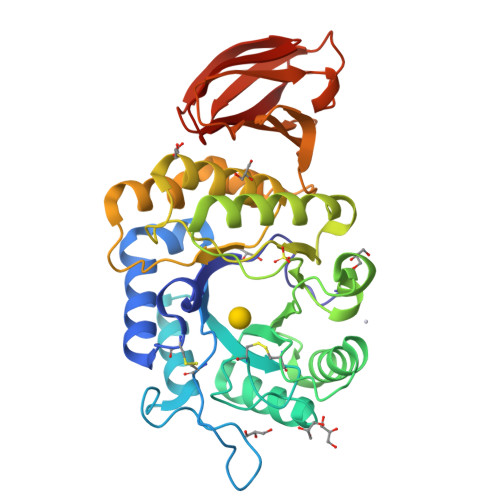
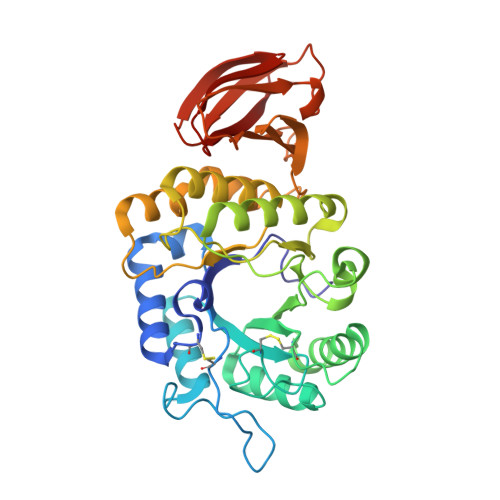
Crystal structure of rice alpha-galactosidase complexed with D-galactose
Fujimoto, Z., Kaneko, S., Momma, M., Kobayashi, H., Mizuno, H.(2003) J Biological Chem 278: 20313-20318
- PubMed: 12657636
- DOI: https://doi.org/10.1074/jbc.M302292200
- Primary Citation of Related Structures:
1UAS - PubMed Abstract:
alpha-Galactosidases catalyze the hydrolysis of alpha-1,6-linked galactosyl residues from galacto-oligosaccharides and polymeric galacto-(gluco)mannans. The crystal structure of rice alpha-galactosidase has been determined at 1.5A resolution using the multiple isomorphous replacement method. The structure consisted of a catalytic domain and a C-terminal domain and was essentially the same as that of alpha-N-acetylgalactosaminidase, which is the same member of glycosyl hydrolase family 27. The catalytic domain had a (beta/alpha)8-barrel structure, and the C-terminal domain was made up of eight beta-strands containing a Greek key motif. The structure was solved as a complex with d-galactose, providing a mode of substrate binding in detail. The d-galactose molecule was found bound in the active site pocket on the C-terminal side of the central beta-barrel of the catalytic domain. The d-galactose molecule consisted of a mixture of two anomers present in a ratio equal to their natural abundance. Structural comparisons of rice alpha-galactosidase with chicken alpha-N-acetylgalactosaminidase provided further understanding of the substrate recognition mechanism in these enzymes.
Organizational Affiliation:
Department of Biochemistry, National Institute of Agrobiological Sciences, Tsukuba, Ibaraki 305-8602, Japan.