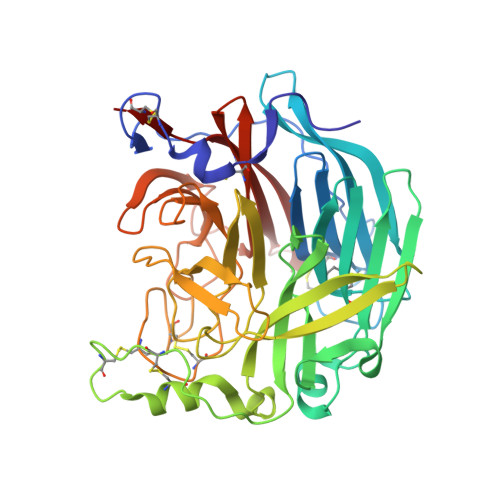
Structure of the Haemagglutinin-neuraminidase from Human Parainfluenza Virus Type III
Lawrence, M.C., Borg, N.A., Streltsov, V.A., Pilling, P.A., Epa, V.C., Varghese, J.N., McKimm-Breschkin, J.L., Colman, P.M.(2004) J Mol Biol 335: 1343-1357
- PubMed: 14729348
- DOI: https://doi.org/10.1016/j.jmb.2003.11.032
- Primary Citation of Related Structures:
1V2I, 1V3B, 1V3C, 1V3D, 1V3E - PubMed Abstract:
The three-dimensional structure of the haemagglutinin-neuraminidase (HN) from a human parainfluenza virus is described at ca 2.0 A resolution, both in native form and in complex with three substrate analogues. In support of earlier work on the structure of the homologous protein from the avian pathogen Newcastle disease virus (NDV), we observe a dimer of beta-propellers and find no evidence for spatially separated sites performing the receptor-binding and neuraminidase functions of the protein. As with the NDV HN, the active site of the HN of parainfluenza viruses is structurally flexible, suggesting that it may be able to switch between a receptor-binding state and a catalytic state. However, in contrast to the NDV structures, we observe no ligand-induced structural changes that extend beyond the active site and modify the dimer interface.
Organizational Affiliation:
CSIRO Health Sciences and Nutrition, 343 Royal Parade, Parkville, Vic 3052, Australia. mike.lawrence@csiro.au