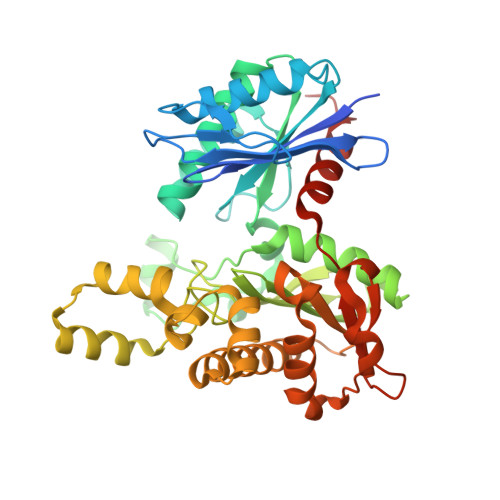
Crystal Structures of ADP and AMPPNP-bound Propionate Kinase (TdcD) from Salmonella typhimurium: Comparison with Members of Acetate and Sugar Kinase/Heat Shock Cognate 70/Actin Superfamily.
Simanshu, D.K., Savithri, H.S., Murthy, M.R.(2005) J Mol Biol 352: 876-892
- PubMed: 16139298
- DOI: https://doi.org/10.1016/j.jmb.2005.07.069
- Primary Citation of Related Structures:
1X3M, 1X3N - PubMed Abstract:
Recently, it has been shown that l-threonine can be catabolized non-oxidatively to propionate via 2-ketobutyrate. Propionate kinase (TdcD; EC 2.7.2.-) catalyses the last step of this metabolic process by enabling the conversion of propionyl phosphate and ADP to propionate and ATP. To provide insights into the substrate-binding pocket and catalytic mechanism of TdcD, the crystal structures of the enzyme from Salmonella typhimurium in complex with ADP and AMPPNP have been determined to resolutions of 2.2A and 2.3A, respectively, by molecular replacement using Methanosarcina thermophila acetate kinase (MAK; EC 2.7.2.1). Propionate kinase, like acetate kinase, contains a fold with the topology betabetabetaalphabetaalphabetaalpha, identical with that of glycerol kinase, hexokinase, heat shock cognaten 70 (Hsc70) and actin, the superfamily of phosphotransferases. The structure consists of two domains with the active site contained in a cleft at the domain interface. Examination of the active site pocket revealed a plausible structural rationale for the greater specificity of the enzyme towards propionate than acetate. This was further confirmed by kinetic studies with the purified enzyme, which showed about ten times lower K(m) for propionate (2.3 mM) than for acetate (26.9 mM). Comparison of TdcD complex structures with those of acetate and sugar kinase/Hsc70/actin obtained with different ligands has permitted the identification of catalytically essential residues involved in substrate binding and catalysis, and points to both structural and mechanistic similarities. In the well-characterized members of this superfamily, ATP phosphoryl transfer or hydrolysis is coupled to a large conformational change in which the two domains close around the active site cleft. The significant amino acid sequence similarity between TdcD and MAK has facilitated study of domain movement, which indicates that the conformation assumed by the two domains in the nucleotide-bound structure of TdcD may represent an intermediate point in the pathway of domain closure.
Organizational Affiliation:
Molecular Biophysics Unit, Indian Institute of Science, Bangalore 560012, India.