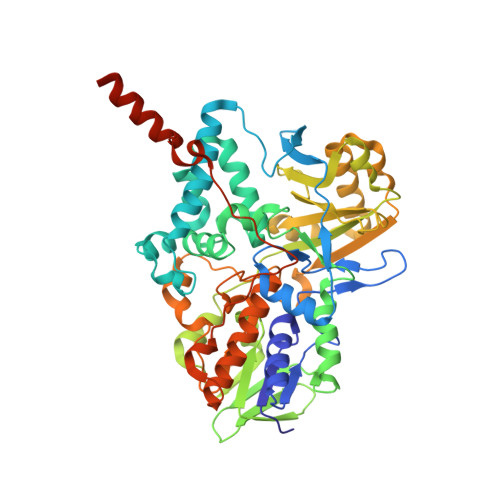
Functional Role of the Aromatic Cage in Human Monoamine Oxidase B: Structures and Catalytic Properties of Tyr435 Mutant Proteins
Li, M., Binda, C., Mattevi, A., Edmondson, D.E.(2006) Biochemistry 45: 4775
- PubMed: 16605246
- DOI: https://doi.org/10.1021/bi051847g
- Primary Citation of Related Structures:
2C70, 2C72, 2C73, 2C75, 2C76 - PubMed Abstract:
Current structural results of several flavin-dependent amine oxidizing enzymes including human monoamine oxidases A and B (MAO A and MAO B) show aromatic amino acid residues oriented approximately perpendicular to the flavin ring, suggesting a functional role in catalysis. In the case of human MAO B, two tyrosyl residues (Y398 and Y435) are found in the substrate binding site on the re face of the covalent flavin ring [Binda et al. (2002) J. Biol. Chem. 277, 23973-23976]. To probe the functional significance of this structure, Tyr435 in MAO B was mutated with the amino acids Phe, His, Leu, or Trp, the mutant proteins expressed in Pichia pastoris, and purified to homogeneity. Each mutant protein contains covalent FAD and exhibits a high level of catalytic functionality. No major alterations in active site structures are detected on comparison of their respective crystal structures with that of WT enzyme. The relative k(cat)/K(m) values for each mutant enzyme show Y435 > Y435F = Y435L = Y435H > Y435W. A similar behavior is also observed with the membrane-bound forms of MAO A and MAO B (MAO A Y444 mutant enzymes are found to be unstable on membrane extraction). p-Nitrobenzylamine is found to be a poor substrate while p-nitrophenethylamine is found to be a good substrate for all WT and mutant forms of MAO B. Analysis of these kinetic and structural data suggests the function of the "aromatic cage" in MAO to include a steric role in substrate binding and access to the flavin coenzyme and to increase the nucleophilicity of the substrate amine moiety. These results are consistent with a proposed polar nucleophilic mechanism for catalytic amine oxidation.
Organizational Affiliation:
Department of Biochemistry, Emory University, 1510 Clifton Road, Atlanta, Georgia 30322, USA.