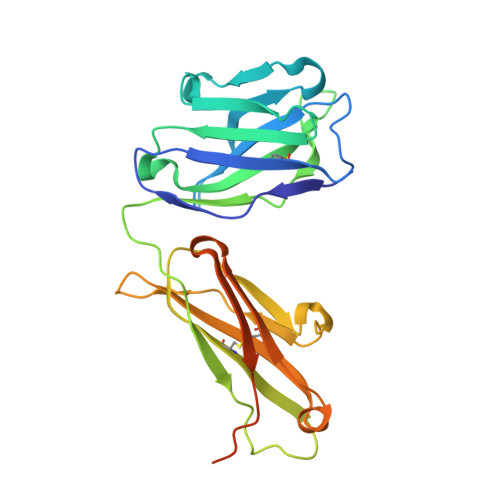
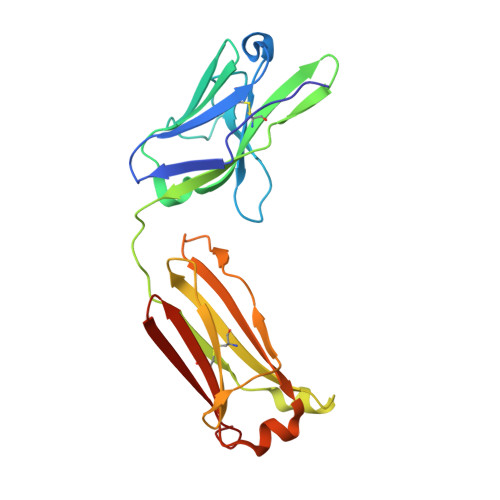
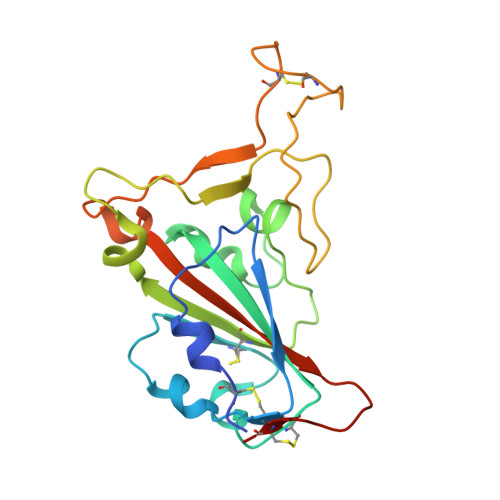
Structure of Severe Acute Respiratory Syndrome Coronavirus Receptor-binding Domain Complexed with Neutralizing Antibody
Prabakaran, P., Gan, J., Feng, Y., Zhu, Z., Choudhry, V., Xiao, X., Ji, X., Dimitrov, D.S.(2006) J Biol Chem 281: 15829-15836
- PubMed: 16597622
- DOI: https://doi.org/10.1074/jbc.M600697200
- Primary Citation of Related Structures:
2DD8, 2G75 - PubMed Abstract:
The severe acute respiratory syndrome coronavirus (SARS-CoV, or SCV), which caused a world-wide epidemic in 2002 and 2003, binds to a receptor, angiotensin-converting enzyme 2 (ACE2), through the receptor-binding domain (RBD) of its envelope (spike, S) glycoprotein. The RBD is very immunogenic; it is a major SCV neutralization determinant and can elicit potent neutralizing antibodies capable of out-competing ACE2. However, the structural basis of RBD immunogenicity, RBD-mediated neutralization, and the role of RBD in entry steps following its binding to ACE2 have not been elucidated. By mimicking immune responses with the use of RBD as an antigen to screen a large human antibody library derived from healthy volunteers, we identified a novel potent cross-reactive SCV-neutralizing monoclonal antibody, m396, which competes with ACE2 for binding to RBD, and determined the crystal structure of the RBD-antibody complex at 2.3-A resolution. The antibody-bound RBD structure is completely defined, revealing two previously unresolved segments (residues 376-381 and 503-512) and a new disulfide bond (between residues 378 and 511). Interestingly, the overall structure of the m396-bound RBD is not significantly different from that of the ACE2-bound RBD. The antibody epitope is dominated by a 10-residue-long protruding beta6-beta7 loop with two putative ACE2-binding hotspot residues (Ile-489 and Tyr-491). These results provide a structural rationale for the function of a major determinant of SCV immunogenicity and neutralization, the development of SCV therapeutics based on the antibody paratope and epitope, and a retrovaccinology approach for the design of anti-SCV vaccines. The available structural information indicates that the SCV entry may not be mediated by ACE2-induced conformational changes in the RBD but may involve other conformational changes or/and yet to be identified coreceptors.
Organizational Affiliation:
Protein Interactions Group, Center for Cancer Research Nanobiology Program, NCI, National Institutes of Health, Frederick, Maryland 21702, USA.