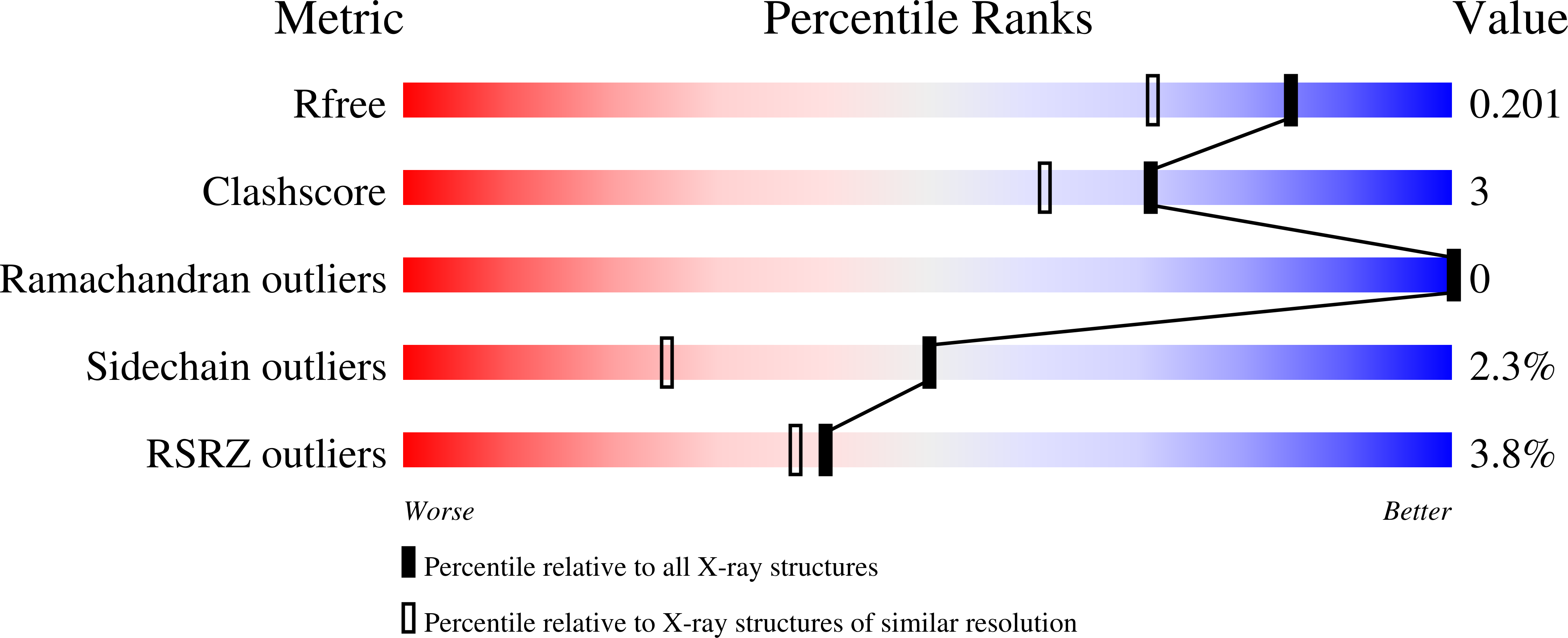
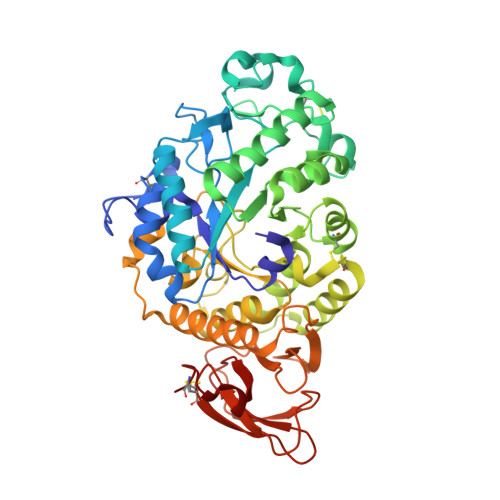
Monoclinic crystal form of Aspergillus niger alpha-amylase in complex with maltose at 1.8 angstroms resolution.
Vujicic-Zagar, A., Dijkstra, B.W.(2006) Acta Crystallogr Sect F Struct Biol Cryst Commun 62: 716-721
- PubMed: 16880540
- DOI: https://doi.org/10.1107/S1744309106024729
- Primary Citation of Related Structures:
2GUY, 2GVY - PubMed Abstract:
Aspergillus niger alpha-amylase catalyses the hydrolysis of alpha-1,4-glucosidic bonds in starch. It shows 100% sequence identity to the A. oryzae homologue (also called TAKA-amylase), three crystal structures of which have been published to date. Two of them belong to the orthorhombic space group P2(1)2(1)2(1) with one molecule per asymmetric unit and one belongs to the monoclinic space group P2(1) with three molecules per asymmetric unit. Here, the purification, crystallization and structure determination of A. niger alpha-amylase crystallized in the monoclinic space group P2(1) with two molecules per asymmetric unit in complex with maltose at 1.8 angstroms resolution is reported. Furthermore, a novel 1.6 angstroms resolution orthorhombic crystal form (space group P2(1)2(1)2) of the native enzyme is presented. Four maltose molecules are observed in the maltose-alpha-amylase complex. Three of these occupy active-site subsites -2 and -1, +1 and +2 and the hitherto unobserved subsites +4 (Asp233, Gly234) and +5 (Asp235). The fourth maltose molecule binds at the distant binding sites d1 (Tyr382) and d2 (Trp385), also previously unobserved. Furthermore, it is shown that the active-site groove permits different binding modes of sugar units at subsites +1 and +2. This flexibility of the active-site cleft close to the catalytic centre might be needed for a productive binding of substrate chains and/or release of products.
Organizational Affiliation:
Laboratory of Biophysical Chemistry, University of Groningen, Nijenborgh 4, 9747AG Groningen, The Netherlands.