Clavulanic Acid Dehydrogenase: Structural and Biochemical Analysis of the Final Step in the Biosynthesis of the Beta-Lactamase Inhibitor Clavulanic Acid
Mackenzie, A.K., Kershaw, N.J., Hernandez, H., Robinson, C.V., Schofield, C.J., Andersson, I.(2007) Biochemistry 46: 1523
- PubMed: 17279617
- DOI: https://doi.org/10.1021/bi061978x
- Primary Citation of Related Structures:
2JAH, 2JAP - PubMed Abstract:
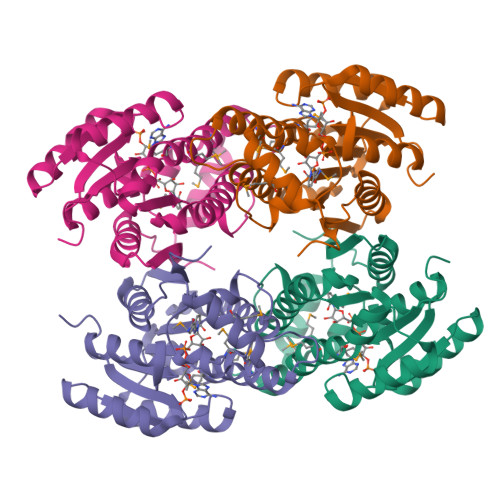
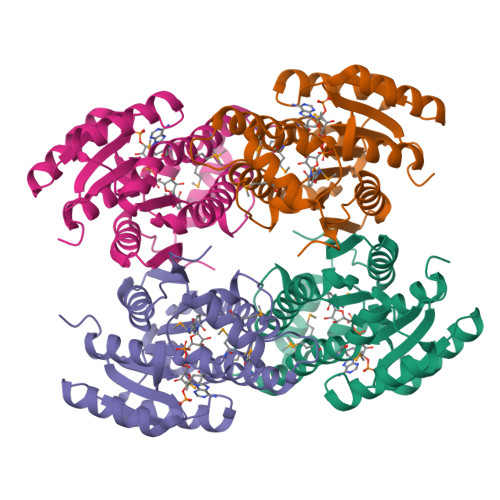
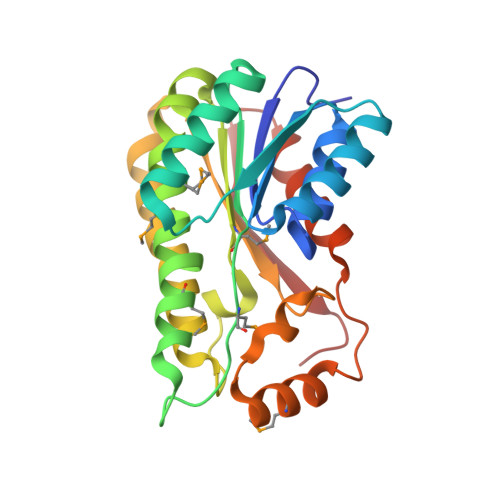
The ultimate step in the biosynthesis of the medicinally important beta-lactamase inhibitor clavulanic acid is catalyzed by clavulanic acid dehydrogenase (CAD). CAD is responsible for the NAPDH-dependent reduction of the unstable intermediate clavulanate-9-aldehyde to yield clavulanic acid. Here, we report biochemical and structural studies on CAD. Biophysical analyses demonstrate that CAD exists as dimeric and tetrameric species in solution. The reaction performed by CAD was shown to be reversible, allowing the use of clavulanic acid for activity analyses. The crystal structure of CAD was solved using single-wavelength anomalous diffraction with a seleno-methionine derivative. The structure reveals that the individual monomers comprise a single domain possessing the Rossmann fold, characteristic of dinucleotide-binding enzymes. The monomers are arranged as tetramers, similar to other tetrameric members of the short-chain dehydrogenase/reductase family. The structure of the unreactive complex of CAD with clavulanic acid and NADPH suggests how CAD is able to catalyze the reduction of clavulanate-9-aldehyde without fragmentation of the bicyclic beta-lactam ring structure. The relative positions of NADPH and clavulanic acid, in the active site, together with the presence of the latter in an eclipsed conformation, rationalizes previous labeling studies demonstrating that the incorporation of the C5 pro-R, but not pro-S, hydrogen of ornithine/arginine into the C9 position of clavulanic acid occurs with overall inversion of configuration.
Organizational Affiliation:
Department of Molecular Biology, Swedish University of Agricultural Sciences, Box 590, S-751 24 Uppsala, Sweden.