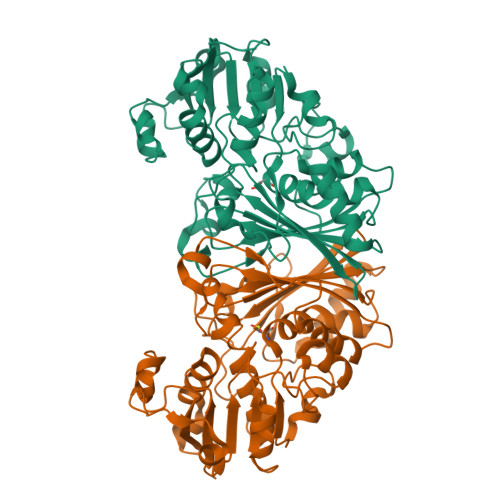
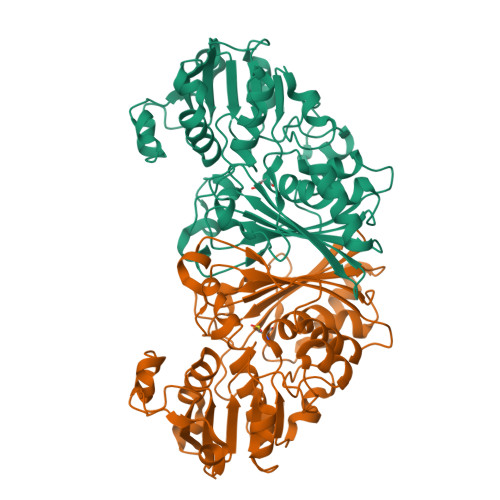
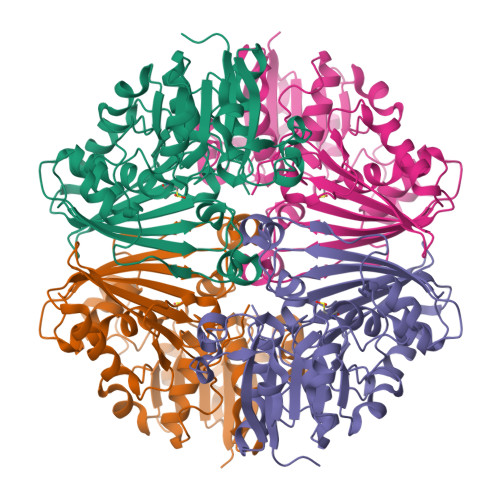
Crystal Structure of N-acetyl-gamma-glutamyl-phosphate Reductase from Mycobacterium tuberculosis in Complex with NADP(+).
Cherney, L.T., Cherney, M.M., Garen, C.R., Niu, C., Moradian, F., James, M.N.(2007) J Mol Biol 367: 1357-1369
- PubMed: 17316682
- DOI: https://doi.org/10.1016/j.jmb.2007.01.033
- Primary Citation of Related Structures:
2I3A, 2I3G, 2NQT - PubMed Abstract:
The enzyme N-acetyl-gamma-glutamyl-phosphate reductase (AGPR) catalyzes the nicotinamide adenine dinucleotide phosphate (NADPH)-dependent reductive dephosphorylation of N-acetyl-gamma-glutamyl-phosphate to N-acetylglutamate-gamma-semialdehyde. This reaction is part of the arginine biosynthetic pathway that is essential for some microorganisms and plants, in particular, for Mycobacterium tuberculosis (Mtb). The structures of apo MtbAGPR in the space groups P2(1)2(1)2(1) and C2 and the structure of MtbAGPR bound to the cofactor NADP(+) have been solved and analyzed. Each MtbAGPR subunit consists of alpha/beta and alpha+beta domains; NADP(+) is bound in the cleft between them. The hydrogen bonds and hydrophobic contacts between the enzyme and cofactor have been examined. Comparison of the apo and the bound enzyme structures has revealed a conformational change in MtbAGPR upon NADP(+) binding. Namely, a loop (Leu88 to His92) moves more than 5 A to confine sterically the cofactor's adenine moiety in a hydrophobic pocket. To identify the catalytically important residues in MtbAGPR, a docking of the substrate to the enzyme has been performed using the present structure of the MtbAGPR/NADP(+) complex. It reveals that residues His217 and His219 could form hydrogen bonds with the docked substrate. In addition, an ion pair could form between the substrate phosphate group and the guanidinium group of Arg114. These interactions optimally place and orient the substrate for subsequent nucleophilic attack by Cys158 on the substrate gamma-carboxyl group. His219 is the most probable general base to accept a proton from Cys158 and an adjacent ion pair interaction with the side-chain carboxyl group of Glu222 could help to stabilize the resulting positive charge on His219. For this catalytic triad to function efficiently it requires a small conformational change of the order of 1 A in the loop containing His217 and His219; this could easily result from the substrate binding.
Organizational Affiliation:
Group in Protein Structure and Function, Department of Biochemistry, University of Alberta, Edmonton, Alberta, Canada T6G 2H7.