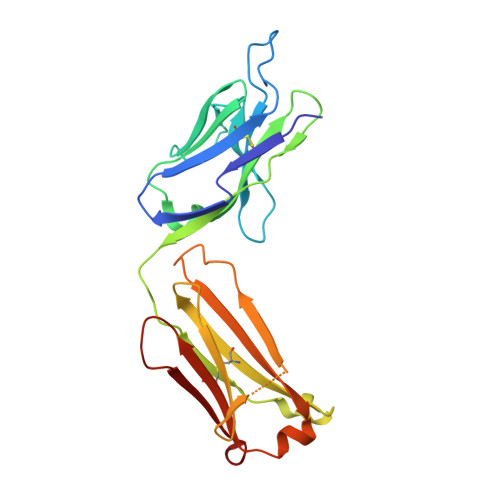
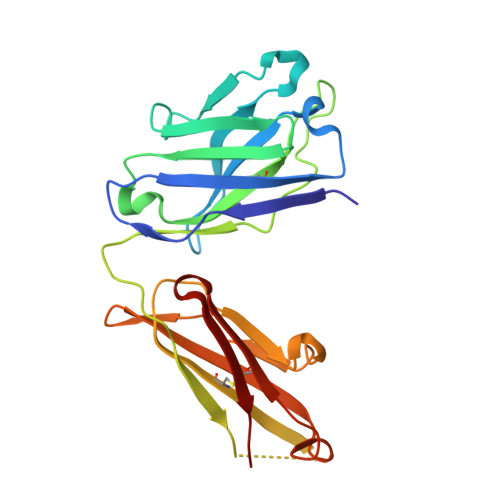
Exploration of specificity in germline monoclonal antibody recognition of a range of natural and synthetic epitopes.
Brooks, C.L., Muller-Loennies, S., Brade, L., Kosma, P., Hirama, T., MacKenzie, C.R., Brade, H., Evans, S.V.(2008) J Mol Biol 377: 450-468
- PubMed: 18272175
- DOI: https://doi.org/10.1016/j.jmb.2008.01.018
- Primary Citation of Related Structures:
2R1W, 2R1X, 2R1Y, 2R23, 2R2B, 2R2E, 2R2H, 3BPC - PubMed Abstract:
To explore the molecular basis of antigen recognition by germline antibodies, we have determined to high resolution the structures of the near-germline monoclonal antibody S25-2 in complex with seven distinct carbohydrate antigens based on the bacterial sugar 3-deoxy-alpha-D-manno-oct-2-ulosonic acid (Kdo). In contrast to previous findings, the inherited germline Kdo monosaccharide binding site is not restricted to this bacterial sugar but is able to accommodate an array of substitutions and chemical modifications of Kdo, including naturally occurring antigens containing the related monosaccharide d-glycero-alpha-d-talo-oct-2-ulosonic acid as well as nonterminal Kdo residues. However, we show by surface plasmon resonance and ELISA how antibody S25-2 specificity is so dependent on the context in which the antigen is presented that a free disaccharide displays strong binding while the same lipid-A-bound disaccharide does not bind. These structures provide insight into how inherited germline genes code for immunoglobulins of limited flexibility that are capable of binding a range of epitopes from which affinity-matured antibodies are generated.
Organizational Affiliation:
Department of Biochemistry and Microbiology, University of Victoria, PO Box 3055 STN CSC, Victoria, BC, Canada.