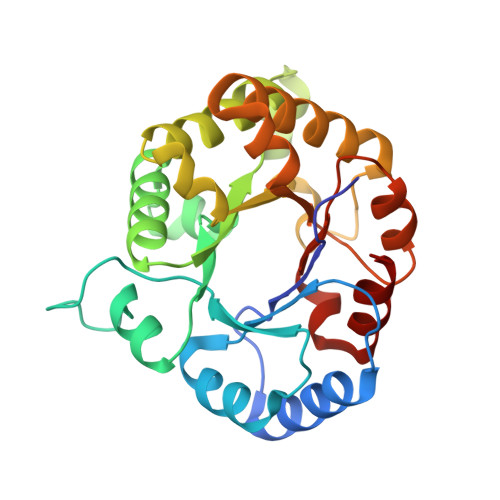
High Resolution Crystal Structures of Triosephosphate Isomerase Complexed with its Suicide Inhibitors: The Conformational Flexibility of the Catalytic Glutamate in its Closed, Liganded Active Site.
Venkatesan, R., Alahuhta, M., Pihko, P.M., Wierenga, R.K.(2011) Protein Sci 20: 1387
- PubMed: 21633986
- DOI: https://doi.org/10.1002/pro.667
- Primary Citation of Related Structures:
2Y61, 2Y62, 2Y63 - PubMed Abstract:
The key residue of the active site of triosephosphate isomerase (TIM) is the catalytic glutamate, which is proposed to be important (i) as a catalytic base, for initiating the reaction, as well as (ii) for the subsequent proton shuttling steps. The structural properties of this glutamate in the liganded complex have been investigated by studying the high resolution crystal structures of typanosomal TIM, complexed with three suicide inhibitors: (S)-glycidol phosphate ((S)-GOP, at 0.99 Å resolution), (R)-glycidol phosphate, ((R)-GOP, at 1.08 Å resolution), and bromohydroxyacetone phosphate (BHAP, at 1.97 Å resolution). The structures show that in the (S)-GOP active site this catalytic glutamate is in the well characterized, competent conformation. However, an unusual side chain conformation is observed in the (R)-GOP and BHAP complexes. In addition, Glu97, salt bridged to the catalytic lysine in the competent active site, adopts an unusual side chain conformation in these two latter complexes. The higher chemical reactivity of (S)-GOP compared with (R)-GOP, as known from solution studies, can be understood: the structures indicate that in the case of (S)-GOP, Glu167 can attack the terminal carbon of the epoxide in a stereoelectronically favored, nearly linear OCO arrangement, but this is not possible for the (R)-GOP isomer. These structures confirm the previously proposed conformational flexibility of the catalytic glutamate in its closed, liganded state. The importance of this conformational flexibility for the proton shuttling steps in the TIM catalytic cycle, which is apparently achieved by a sliding motion of the side chain carboxylate group above the enediolate plane, is also discussed.
Organizational Affiliation:
Biocenter Oulu and Department of Biochemistry, University of Oulu, Finland.