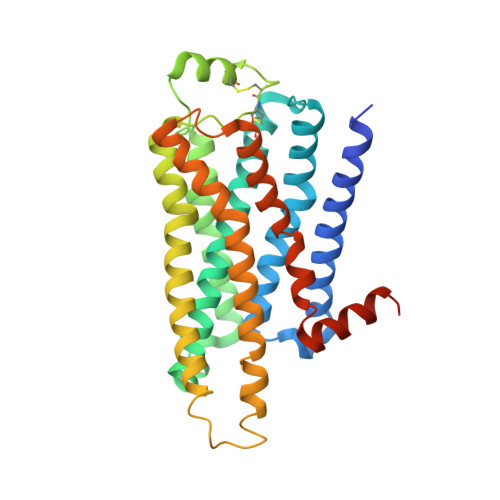
Two Distinct Conformations of Helix 6 Observed in Antagonist-Bound Structures of a {Beta}1- Adrenergic Receptor.
Moukhametzianov, R., Warne, T., Edwards, P.C., Serrano-Vega, M.J., Leslie, A.G., Tate, C.G., Schertler, G.F.(2011) Proc Natl Acad Sci U S A 108: 8228
- PubMed: 21540331
- DOI: https://doi.org/10.1073/pnas.1100185108
- Primary Citation of Related Structures:
2YCW, 2YCX, 2YCY, 2YCZ - PubMed Abstract:
The β(1)-adrenergic receptor (β(1)AR) is a G-protein-coupled receptor whose inactive state structure was determined using a thermostabilized mutant (β(1)AR-M23). However, it was not thought to be in a fully inactivated state because there was no salt bridge between Arg139 and Glu285 linking the cytoplasmic ends of transmembrane helices 3 and 6 (the R(3.50) - D/E(6.30) "ionic lock"). Here we compare eight new structures of β(1)AR-M23, determined from crystallographically independent molecules in four different crystals with three different antagonists bound. These structures are all in the inactive R state and show clear electron density for cytoplasmic loop 3 linking transmembrane helices 5 and 6 that had not been seen previously. Despite significantly different crystal packing interactions, there are only two distinct conformations of the cytoplasmic end of helix 6, bent and straight. In the bent conformation, the Arg139-Glu285 salt bridge is present, as in the crystal structure of dark-state rhodopsin. The straight conformation, observed in previously solved structures of β-receptors, results in the ends of helices 3 and 6 being too far apart for the ionic lock to form. In the bent conformation, the R(3.50)-E(6.30) distance is significantly longer than in rhodopsin, suggesting that the interaction is also weaker, which could explain the high basal activity in β(1)AR compared to rhodopsin. Many mutations that increase the constitutive activity of G-protein-coupled receptors are found in the bent region at the cytoplasmic end of helix 6, supporting the idea that this region plays an important role in receptor activation.
Organizational Affiliation:
Medical Research Council Laboratory of Molecular Biology, Hills Road, Cambridge CB2 0QH, United Kingdom.