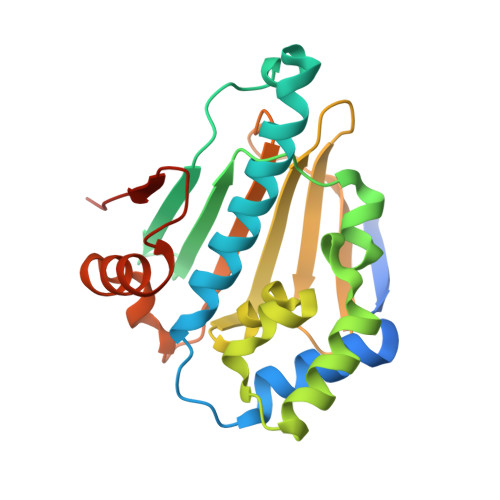
Co-Crystalization and in Vitro Biological Characterization of 5-Aryl-4-(5-Substituted-2-4-Dihydroxyphenyl)-1,2,3-Thiadiazole Hsp90 Inhibitors.
Sharp, S.Y., Roe, S.M., Kazlauskas, E., Cikotiene, I., Workman, P., Matulis, D., Prodromou, C.(2012) PLoS One 7: 44642
- PubMed: 22984537
- DOI: https://doi.org/10.1371/journal.pone.0044642
- Primary Citation of Related Structures:
2YI0, 2YI5, 2YI6, 2YI7 - PubMed Abstract:
A potential therapeutic strategy for targeting cancer that has gained much interest is the inhibition of the ATP binding and ATPase activity of the molecular chaperone Hsp90. We have determined the structure of the human Hsp90α N-terminal domain in complex with a series of 5-aryl-4-(5-substituted-2-4-dihydroxyphenyl)-1,2,3-thiadiazoles. The structures provide the molecular details for the activity of these inhibitors. One of these inhibitors, ICPD 34, causes a structural change that affects a mobile loop, which adopts a conformation similar to that seen in complexes with ADP, rather than the conformation generally seen with the pyrazole/isoxazole-resorcinol class of inhibitors. Competitive binding to the Hsp90 N-terminal domain was observed in a biochemical assay, and these compounds showed antiproliferative activity and induced apoptosis in the HCT116 human colon cancer cell line. These inhibitors also caused induction of the heat shock response with the upregulation of Hsp72 and Hsp27 protein expression and the depletion of Hsp90 clients, CRAF, ERBB2 and CDK4, thus confirming that antiproliferative activity was through the inhibition of Hsp90. The presence of increased levels of the cleavage product of PARP indicated apoptosis in response to Hsp90 inhibitors. This work provides a framework for the further optimization of thiadiazole inhibitors of Hsp90. Importantly, we demonstrate that the thiadiazole inhibitors display a more limited core set of interactions relative to the clinical trial candidate NVP-AUY922, and consequently may be less susceptible to resistance derived through mutations in Hsp90.
Organizational Affiliation:
Cancer Research UK Cancer Therapeutics Unit, Division of Cancer Therapeutics, The Institute of Cancer Research, Haddow Laboratories, Sutton, Surrey, United Kingdom.