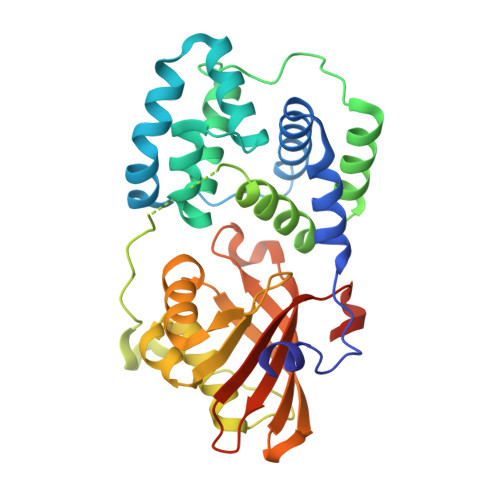
Structural determinants underlying photoprotection in the photoactive orange carotenoid protein of cyanobacteria.
Wilson, A., Kinney, J.N., Zwart, P.H., Punginelli, C., D'Haene, S., Perreau, F., Klein, M.G., Kirilovsky, D., Kerfeld, C.A.(2010) J Biol Chem 285: 18364-18375
- PubMed: 20368334
- DOI: https://doi.org/10.1074/jbc.M110.115709
- Primary Citation of Related Structures:
3MG1, 3MG2, 3MG3 - PubMed Abstract:
The photoprotective processes of photosynthetic organisms involve the dissipation of excess absorbed light energy as heat. Photoprotection in cyanobacteria is mechanistically distinct from that in plants; it involves the orange carotenoid protein (OCP), a water-soluble protein containing a single carotenoid. The OCP is a new member of the family of blue light-photoactive proteins; blue-green light triggers the OCP-mediated photoprotective response. Here we report structural and functional characterization of the wild type and two mutant forms of the OCP, from the model organism Synechocystis PCC6803. The structural analysis provides high resolution detail of the carotenoid-protein interactions that underlie the optical properties of the OCP, unique among carotenoid-proteins in binding a single pigment per polypeptide chain. Collectively, these data implicate several key amino acids in the function of the OCP and reveal that the photoconversion and photoprotective responses of the OCP to blue-green light can be decoupled.
Organizational Affiliation:
Commissariat à l'Energie Atomique, Institut de Biologie et Technologies de Saclay, CNRS, URA 2906, 91191 Gif sur Yvette, France.