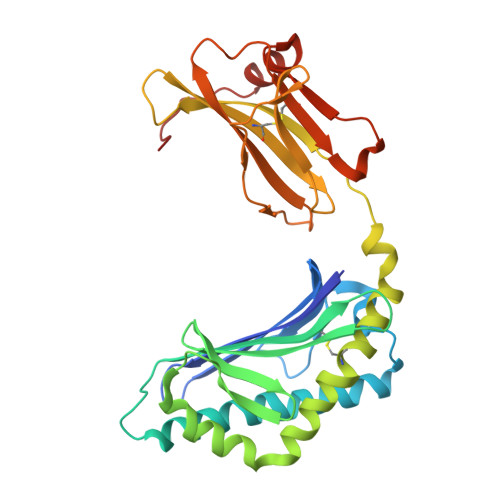
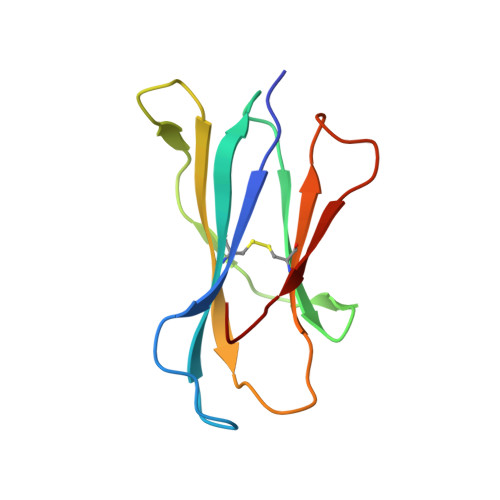
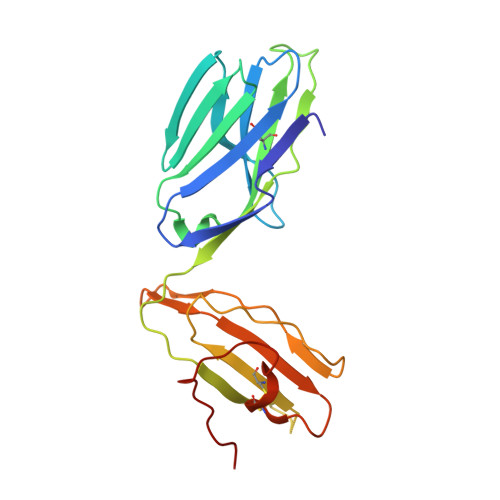
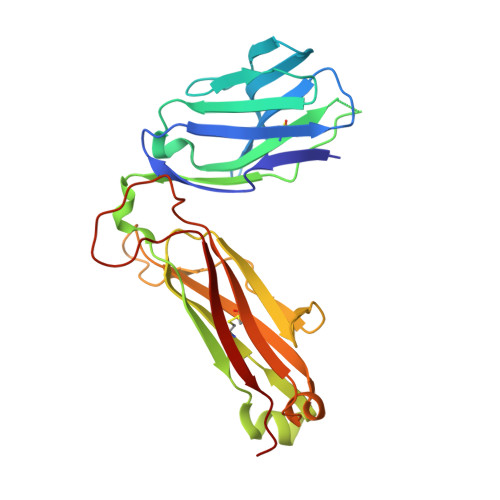
NKT TCR Recognition of CD1d-{alpha}-C-Galactosylceramide.
Patel, O., Cameron, G., Pellicci, D.G., Liu, Z., Byun, H.S., Beddoe, T., McCluskey, J., Franck, R.W., Castano, A.R., Harrak, Y., Llebaria, A., Bittman, R., Porcelli, S.A., Godfrey, D.I., Rossjohn, J.(2011) J Immunol 187: 4705-4713
- PubMed: 21964029
- DOI: https://doi.org/10.4049/jimmunol.1100794
- Primary Citation of Related Structures:
3TN0 - PubMed Abstract:
NKT cells respond to a variety of CD1d-restricted glycolipid Ags that are structurally related to the prototypic Ag α-galactosylceramide (α-GalCer). A modified analog of α-GalCer with a carbon-based glycosidic linkage (α-C-GalCer) has generated great interest because of its apparent ability to promote prolonged, Th1-biased immune responses. In this study, we report the activation of spleen NKT cells to α-C-GalCer, and related C-glycoside ligands, is weaker than that of α-GalCer. Furthermore, the Vβ8.2 and Vβ7 NKT TCR affinity for CD1d-α-C-GalCer, and some related analogs, is ∼10-fold lower than that for the NKT TCR-CD1d-α-GalCer interaction. Nevertheless, the crystal structure of the Vβ8.2 NKT TCR-CD1d-α-C-GalCer complex is similar to that of the corresponding NKT TCR-CD1d-α-GalCer complex, although subtle differences at the interface provide a basis for understanding the lower affinity of the NKT TCR-CD1d-α-C-GalCer interaction. Our findings support the concept that for CD1d-restricted NKT cells, altered glycolipid ligands can promote markedly different responses while adopting similar TCR-docking topologies.
Organizational Affiliation:
Australian Research Council Centre of Excellence in Structural and Functional Microbial Genomics, Department of Biochemistry and Molecular Biology, School of Biomedical Sciences, Monash University, Clayton, Victoria 3800, Australia.