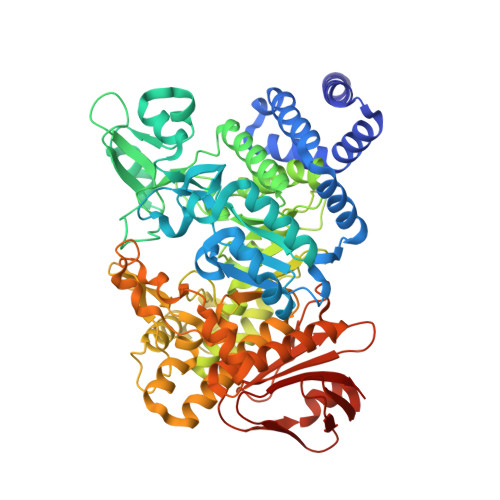
Structural Investigation of the Thermostability and Product Specificity of Amylosucrase from the Bacterium Deinococcus geothermalis.
Guerin, F., Barbe, S., Pizzut-Serin, S., Potocki-Veronese, G., Guieysse, D., Guillet, V., Monsan, P., Mourey, L., Remaud-Simeon, M., Andre, I., Tranier, S.(2012) J Biol Chem 287: 6642-6654
- PubMed: 22210773
- DOI: https://doi.org/10.1074/jbc.M111.322917
- Primary Citation of Related Structures:
3UCQ, 3UEQ, 3UER - PubMed Abstract:
Amylosucrases are sucrose-utilizing α-transglucosidases that naturally catalyze the synthesis of α-glucans, linked exclusively through α1,4-linkages. Side products and in particular sucrose isomers such as turanose and trehalulose are also produced by these enzymes. Here, we report the first structural and biophysical characterization of the most thermostable amylosucrase identified so far, the amylosucrase from Deinoccocus geothermalis (DgAS). The three-dimensional structure revealed a homodimeric quaternary organization, never reported before for other amylosucrases. A sequence signature of dimerization was identified from the analysis of the dimer interface and sequence alignments. By rigidifying the DgAS structure, the quaternary organization is likely to participate in the enhanced thermal stability of the protein. Amylosucrase specificity with respect to sucrose isomer formation (turanose or trehalulose) was also investigated. We report the first structures of the amylosucrases from Deinococcus geothermalis and Neisseria polysaccharea in complex with turanose. In the amylosucrase from N. polysaccharea (NpAS), key residues were found to force the fructosyl moiety to bind in an open state with the O3' ideally positioned to explain the preferential formation of turanose by NpAS. Such residues are either not present or not similarly placed in DgAS. As a consequence, DgAS binds the furanoid tautomers of fructose through a weak network of interactions to enable turanose formation. Such topology at subsite +1 is likely favoring other possible fructose binding modes in agreement with the higher amount of trehalulose formed by DgAS. Our findings help to understand the inter-relationships between amylosucrase structure, flexibility, function, and stability and provide new insight for amylosucrase design.
Organizational Affiliation:
Université de Toulouse; INSA, UPS, INP, LISBP, 135 Avenue de Rangueil, F-31077 Toulouse, France.